Available via license: CC BY 4.0
Content may be subject to copyright.
Citation: Feng, T.; Liu, Q.; Xu, Z.-Y.;
Li, H.-T.; Wei, W.; Shi, R.-C.; Zhang,
L.; Cao, Y.-M.; Liu, S.-Z. Design,
Synthesis, Herbicidal Activity, and
Structure–Activity Relationship
Study of Novel 6-(5-Aryl-Substituted-
1-Pyrazolyl)-2-Picolinic Acid as
Potential Herbicides. Molecules 2023,
28, 1431. https://doi.org/10.3390/
molecules28031431
Academic Editor: Łukasz
Chrzanowski
Received: 27 December 2022
Revised: 18 January 2023
Accepted: 30 January 2023
Published: 2 February 2023
Copyright: © 2023 by the authors.
Licensee MDPI, Basel, Switzerland.
This article is an open access article
distributed under the terms and
conditions of the Creative Commons
Attribution (CC BY) license (https://
creativecommons.org/licenses/by/
4.0/).
molecules
Article
Design, Synthesis, Herbicidal Activity, and Structure–Activity
Relationship Study of Novel 6-(5-Aryl-Substituted-1-Pyrazolyl)-
2-Picolinic Acid as Potential Herbicides
Tong Feng 1,2, Qing Liu 1,2, Zhi-Yuan Xu 1,2, Hui-Ting Li 1, Wei Wei 1, Rong-Chuan Shi 1, Li Zhang 1,2 ,
Yi-Ming Cao 1,2 ,* and Shang-Zhong Liu 1 ,2 ,*
1Innovation Center of Pesticide Research, Department of Applied Chemistry, College of Science,
China Agricultural University, Beijing 100193, China
2Key Laboratory of National Forestry and Grassland Administration on Pest Chemical Control,
China Agricultural University, Beijing 100193, China
*
Correspondence: caoym@cau.edu.cn (Y.-M.C.); shangzho@cau.edu.cn (S.-Z.L.); Tel.: +86-10-62731070 (S.-Z.L.)
Abstract:
Picolinic acid and picolinate compounds are a remarkable class of synthetic auxin herbi-
cides. In recent years, two new picolinate compounds, halauxifen-methyl (Arylex
TM
active) and
florpyrauxifen-benzyl (Rinskor
TM
active), have been launched as novel herbicides. Using their struc-
tural skeleton as a template, 33 4-amino-3,5-dicholor-6-(5-aryl-substituted-1-pytazolyl)-2-picolinic
acid compounds were designed and synthesized for the discovery of compounds with potent herbi-
cidal activity. The compounds were tested for inhibitory activity against the growth of Arabidopsis
thaliana roots, and the results demonstrated that the IC
50
value of compound
V-7
was 45 times lower
than that of the halauxifen-methyl commercial herbicide. Molecular docking analyses revealed that
compound
V-7
docked with the receptor auxin-signaling F-box protein 5 (AFB5) more intensively
than picloram. An adaptive three-dimensional quantitative structure–activity relationship model
was constructed from these IC
50
values to guide the next step of the synthetic strategy. Herbicidal
tests of the new compounds indicated that compound
V-8
exhibited better post-emergence herbicidal
activity than picloram at a dosage of 300 gha
−1
, and it was also safe for corn, wheat, and sorghum at
this dosage. These results demonstrated that 6-(5-aryl-substituted-1-pyrazolyl)-2-picolinic acid com-
pounds could be used as potential lead structures in the discovery of novel synthetic
auxin herbicides
.
Keywords: AFB5; docking; picolinic acid; synthesis; synthetic auxin herbicides; 3D-QSAR
1. Introduction
The world’s population is continuing to grow and is expected to reach 8 billion in
November 2022 [
1
]. Demand for food is also increasing but arable land increase is far
below the need of food from population increase [
2
]. Ensuring the unit production of
plants conducting photosynthesis in agricultural practice is one of the key measures to
meet the food demand [3]. The feeding and growth of agriculture pests in cultivated land
ecological systems could harm crop growth and reduce crop productivity. For instance,
weeds compete with crops for light, water, and nutrients, and influence crop growth and
productivity. Several measures have been taken to combat agricultural pests, such as
insect pests, fungi, and weeds; among them, chemical control is the most economic and
effective method. Synthetic herbicides play an important role in weed control and crop
yield enhancement; however, the large-scale and long-term application of some herbicides
compels weeds to generate resistance, which requires the continuous discovery of new
herbicidal molecules with low resistance, low toxicity, and high efficiency [
4
]. Synthetic
auxin herbicides with structures of phenoxyacetic acid, benzoic acid, pyridinoxyacetic
acid, pyridinecarboxylic acid, and 6-aryl-2-pyridinecarboxylate are important chemical
herbicides; from the international herbicide-resistant weed database [
5
], the number of
Molecules 2023,28, 1431. https://doi.org/10.3390/molecules28031431 https://www.mdpi.com/journal/molecules
Molecules 2023,28, 1431 2 of 20
weed species that are resistant to synthetic auxin herbicides is significantly increasing at
a slower rate than others [
6
] because of their unique mode of action and specific binding
sites in target proteins, indicating that they have great potential for the development
of new herbicides. In 2007, Tan et al. presented the crystal structure of the Arabidopsis
transport inhibitor response 1 (TIR1)-ASK1 complex and established the first structural
model of a plant hormone receptor and the mode of binding between auxins and a target
protein [
7
]. Their study guides researchers in exploring the mode of action of synthetic
auxin compounds and designing new computer-aided auxin molecules. Studies have
reported that 2-picolinic acid synthetic auxins herbicides have physiological functions
similar to those of IAA, 2,4-D, and other auxin analogs [
8
,
9
]; however, they bind to AFB5
rather than TIR1, which is a binding protein of IAA [9–12].
Picloram and clopyralid were commercialized as herbicides in the 1960s and 1975, at
application rates of 125–1120 and 105–500 gha
−1
, respectively. Subsequently, aminopy-
ralid, discovered by modifying picloram, was commercialized as a herbicide in 2006 with
application rates of 5–120 gha
−1
[
13
]. In 2015, Jeffrey B. Epp et al. reported that 6-aryl-
2-picolinates exhibit excellent herbicidal activities by replacing the chlorine atom with a
phenyl group at position 6 of 2-picolinic acid herbicides, and they discovered two novel
picolinate herbicides, halauxifen-methyl and florpyrauxifen-benzyl [
13
–
15
]. Even though
these herbicides act on complex auxin-binding proteins, weeds inevitably generate resis-
tance with long-term extensive application; for instance, some weeds were observed to be
resistant to picloram [
16
,
17
]. In this work, we attempted to modify the chemical structure
of picloram to obtain highly effective herbicidal molecules.
In 2021, Yang et al. obtained 3-chloro-6-pyrazolyl-2-picolinic acids and their ester
derivatives by modifying clopyralid using substituted pyrazole rings [
18
]. Bioassay tests
indicated that compound
c5
exhibited better postemergence herbicidal activity and broader
herbicidal activity at a dosage of 400 gha
−1
than clopyralid. This indicates that the in-
troduction of pyrazolyl at position 6 of 2-picolinic acid could be a potential strategy for
discovering a novel synthetic auxin herbicide (Figure 1).
Molecules 2023, 28, x FOR PEER REVIEW 2 of 21
compels weeds to generate resistance, which requires the continuous discovery of new
herbicidal molecules with low resistance, low toxicity, and high efficiency [4]. Synthetic
auxin herbicides with structures of phenoxyacetic acid, benzoic acid, pyridinoxyacetic
acid, pyridinecarboxylic acid, and 6-aryl-2-pyridinecarboxylate are important chemical
herbicides; from the international herbicide-resistant weed database [5], the number of
weed species that are resistant to synthetic auxin herbicides is significantly increasing at
a slower rate than others [6] because of their unique mode of action and specific binding
sites in target proteins, indicating that they have great potential for the development of
new herbicides. In 2007, Tan et al. presented the crystal structure of the Arabidopsis
transport inhibitor response 1 (TIR1)-ASK1 complex and established the first structural
model of a plant hormone receptor and the mode of binding between auxins and a target
protein [7]. Their study guides researchers in exploring the mode of action of synthetic
auxin compounds and designing new computer-aided auxin molecules. Studies have re-
ported that 2-picolinic acid synthetic auxins herbicides have physiological functions sim-
ilar to those of IAA, 2,4-D, and other auxin analogs [8,9]; however, they bind to AFB5
rather than TIR1, which is a binding protein of IAA [9–12].
Picloram and clopyralid were commercialized as herbicides in the 1960s and 1975, at
application rates of 125–1120 and 105–500 gha−1, respectively. Subsequently, aminopyra-
lid, discovered by modifying picloram, was commercialized as a herbicide in 2006 with
application rates of 5–120 gha−1 [13]. In 2015, Jeffrey B. Epp et al. reported that 6-aryl-2-
picolinates exhibit excellent herbicidal activities by replacing the chlorine atom with a
phenyl group at position 6 of 2-picolinic acid herbicides, and they discovered two novel
picolinate herbicides, halauxifen-methyl and florpyrauxifen-benzyl [13–15]. Even though
these herbicides act on complex auxin-binding proteins, weeds inevitably generate re-
sistance with long-term extensive application; for instance, some weeds were observed to
be resistant to picloram [16,17]. In this work, we attempted to modify the chemical struc-
ture of picloram to obtain highly effective herbicidal molecules.
In 2021, Yang et al. obtained 3-chloro-6-pyrazolyl-2-picolinic acids and their ester de-
rivatives by modifying clopyralid using substituted pyrazole rings [18]. Bioassay tests in-
dicated that compound c5 exhibited better postemergence herbicidal activity and broader
herbicidal activity at a dosage of 400 g ha−1 than clopyralid. This indicates that the intro-
duction of pyrazolyl at position 6 of 2-picolinic acid could be a potential strategy for dis-
covering a novel synthetic auxin herbicide (Figure 1).
Figure 1. Structures of picolinic acid and 6-aryl-2-picolinate synthetic auxin herbicides. The refer-
ences involved include [13–18].
Figure 1.
Structures of picolinic acid and 6-aryl-2-picolinate synthetic auxin herbicides. The references
involved include [13–18].
Molecules 2023,28, 1431 3 of 20
Pyrazoles are aromatic five-membered heterocyclicring molecules, and their structural
characteristics have garnered considerable attention among researchers for the incorpo-
ration of pyrazolyl with different substituents into various structures. Therefore, they
exist in a large number of biologically active molecules relevant to the pharmaceutical
and agrochemical industries [
19
]. To date, some molecules containing pyrazole have been
launched as herbicides, such as benzofenap, pyrazoxyfen, and cypyrafluone. Meanwhile,
molecules containing pyrazole have exhibited potential bioactivity in recent research and
patents [20–23] (Figure 2).
Molecules 2023, 28, x FOR PEER REVIEW 3 of 21
Pyrazoles are aromatic five-membered heterocyclic ring molecules, and their struc-
tural characteristics have garnered considerable attention among researchers for the in-
corporation of pyrazolyl with different substituents into various structures. Therefore,
they exist in a large number of biologically active molecules relevant to the pharmaceuti-
cal and agrochemical industries [19]. To date, some molecules containing pyrazole have
been launched as herbicides, such as benzofenap, pyrazoxyfen, and cypyrafluone. Mean-
while, molecules containing pyrazole have exhibited potential bioactivity in recent re-
search and patents [20–23] (Figure 2).
Figure 2. Structures of commercial herbicides containing pyrazoles.
Inspired by the discovery of 6-aryl-2-picolinate herbicides, we designed and synthe-
sized 33 4-amino-3,5-dichloro-6-pyrazolyl-2-picolinic acids with a phenyl-substituted py-
razole replacing the chlorine atom at position 6 of picloram, in order to explore a new
herbicidal molecule (Figure 3). The inhibition of Arabidopsis thaliana root growth, herbi-
cidal activities, and crop selectivity were tested, and the quantitative structure–activity
relationship (QSAR), molecular docking, and mode of action were also preliminarily ex-
plored.
Figure 3. Design strategy of target compounds.
2. Results and Discussion
2.1. Chemistry
The general synthetic procedure is illustrated in Scheme 1. All materials were com-
mercially available. Intermediate II was prepared via a nucleophilic substitution reaction
in which the chlorine atom at position 6 of picloram was replaced by hydrazine hydrate
[24]. Intermediate III was obtained via a Claisen reaction between ethyl acetate or ethyl
di/trifluoroacetate and methyl ketones [25]. Moreover, intermediate IV was synthesized
via the Knorr cyclization reaction of intermediate II and intermediate III [26]. Owing to
the unsymmetrical intermediate III, two region isomers are often obtained in this reaction,
and 5-aryl-pyrazolyl substituted product is 10 times more abundant than 3-aryl-pyrazolyl
substituted product when R1 is an electron-withdrawing group (R1 = CHF2; and CF3). A
possible reason for this is that the carbon atom on the carbonyl group connecting R1 is a
more deficient electron and is more attractive to the nitrogen atom containing the lone
electron pair in 6-hydrazinyl-2-picolinitrile. When R1 was a non-substituted alkyl group
(R1 = Me), the regio-selectivity weakened and the ratio of the 5-aryl-pyrazolyl substituted
product to 3-aryl-pyrazolyl substituted product was in the range 3:1–5:1. Finally, the cy-
ano group in intermediate IV was hydrolyzed as a carboxylic acid group to yield the tar-
get compound V [27]. All the target compounds were characterized through HRMS and
NMR, and their NMR spectra and data are shown in the Supplementary Materials.
Figure 2. Structures of commercial herbicides containing pyrazoles.
Inspired by the discovery of 6-aryl-2-picolinate herbicides, we designed and syn-
thesized 33 4-amino-3,5-dichloro-6-pyrazolyl-2-picolinic acids with a phenyl-substituted
pyrazole replacing the chlorine atom at position 6 of picloram, in order to explore a new her-
bicidal molecule (Figure 3). The inhibition of Arabidopsis thaliana root growth, herbicidal
activities, and crop selectivity were tested, and the quantitative structure–activity relation-
ship (QSAR), molecular docking, and mode of action were also preliminarily explored.
Molecules 2023, 28, x FOR PEER REVIEW 3 of 21
Pyrazoles are aromatic five-membered heterocyclic ring molecules, and their struc-
tural characteristics have garnered considerable attention among researchers for the in-
corporation of pyrazolyl with different substituents into various structures. Therefore,
they exist in a large number of biologically active molecules relevant to the pharmaceuti-
cal and agrochemical industries [19]. To date, some molecules containing pyrazole have
been launched as herbicides, such as benzofenap, pyrazoxyfen, and cypyrafluone. Mean-
while, molecules containing pyrazole have exhibited potential bioactivity in recent re-
search and patents [20–23] (Figure 2).
Figure 2. Structures of commercial herbicides containing pyrazoles.
Inspired by the discovery of 6-aryl-2-picolinate herbicides, we designed and synthe-
sized 33 4-amino-3,5-dichloro-6-pyrazolyl-2-picolinic acids with a phenyl-substituted py-
razole replacing the chlorine atom at position 6 of picloram, in order to explore a new
herbicidal molecule (Figure 3). The inhibition of Arabidopsis thaliana root growth, herbi-
cidal activities, and crop selectivity were tested, and the quantitative structure–activity
relationship (QSAR), molecular docking, and mode of action were also preliminarily ex-
plored.
Figure 3. Design strategy of target compounds.
2. Results and Discussion
2.1. Chemistry
The general synthetic procedure is illustrated in Scheme 1. All materials were com-
mercially available. Intermediate II was prepared via a nucleophilic substitution reaction
in which the chlorine atom at position 6 of picloram was replaced by hydrazine hydrate
[24]. Intermediate III was obtained via a Claisen reaction between ethyl acetate or ethyl
di/trifluoroacetate and methyl ketones [25]. Moreover, intermediate IV was synthesized
via the Knorr cyclization reaction of intermediate II and intermediate III [26]. Owing to
the unsymmetrical intermediate III, two region isomers are often obtained in this reaction,
and 5-aryl-pyrazolyl substituted product is 10 times more abundant than 3-aryl-pyrazolyl
substituted product when R1 is an electron-withdrawing group (R1 = CHF2; and CF3). A
possible reason for this is that the carbon atom on the carbonyl group connecting R1 is a
more deficient electron and is more attractive to the nitrogen atom containing the lone
electron pair in 6-hydrazinyl-2-picolinitrile. When R1 was a non-substituted alkyl group
(R1 = Me), the regio-selectivity weakened and the ratio of the 5-aryl-pyrazolyl substituted
product to 3-aryl-pyrazolyl substituted product was in the range 3:1–5:1. Finally, the cy-
ano group in intermediate IV was hydrolyzed as a carboxylic acid group to yield the tar-
get compound V [27]. All the target compounds were characterized through HRMS and
NMR, and their NMR spectra and data are shown in the Supplementary Materials.
Figure 3. Design strategy of target compounds.
2. Results and Discussion
2.1. Chemistry
The general synthetic procedure is illustrated in Scheme 1. All materials were com-
mercially available. Intermediate
II
was prepared via a nucleophilic substitution reac-
tion in which the chlorine atom at position 6 of picloram was replaced by hydrazine
hydrate [
24
]. Intermediate
III
was obtained via a Claisen reaction between ethyl acetate
or ethyl di/trifluoroacetate and methyl ketones [
25
]. Moreover, intermediate
IV
was syn-
thesized via the Knorr cyclization reaction of intermediate
II
and intermediate
III [26]
.
Owing to the unsymmetrical intermediate
III
, two region isomers are often obtained in this
reaction, and 5-aryl-pyrazolyl substituted product is 10 times more abundant than 3-aryl-
pyrazolyl substituted product when R
1
is an electron-withdrawing group (R
1
= CHF
2
; and
CF
3
). A possible reason for this is that the carbon atom on the carbonyl group connecting R
1
is a more deficient electron and is more attractive to the nitrogen atom containing the lone
electron pair in 6-hydrazinyl-2-picolinitrile. When R
1
was a non-substituted alkyl group
(R
1
= Me), the regio-selectivity weakened and the ratio of the 5-aryl-pyrazolyl substituted
product to 3-aryl-pyrazolyl substituted product was in the range 3:1–5:1. Finally, the cyano
group in intermediate
IV
was hydrolyzed as a carboxylic acid group to yield the target
compound
V [27]
. All the target compounds were characterized through HRMS and NMR,
and their NMR spectra and data are shown in the Supplementary Materials.
Molecules 2023,28, 1431 4 of 20
Molecules 2023, 28, x FOR PEER REVIEW 4 of 21
V-1: R1 = Me, R2 = 4-Me
V-2: R1 = CHF2, R2 = 4-Me
V-3: R1 = CF3, R2 = 4-Me
V-4: R1 = Me, R2 = 4-F
V-5: R1 = CHF2, R2 = 4-F
V-6: R1 = CF3, R2 = 4-F
V-7: R1 = Me, R2 = 4-Cl
V-8: R1 = CHF2, R2 = 4-Cl
V-9: R1 = CF3, R2 = 4-Cl
V-10: R1 = Me, R2 = 3-Cl
V-11: R1 = CHF2, R2 = 3-Cl
V-12: R1 = CF3, R2 = 3-Cl
V-13: R1 = Me, R2 = 4-Br
V-14: R1 = CHF2, R2 = 4-Br
V-15: R1 = CF3, R2 = 4-Br
V-16: R1 = Me, R2 = 2-Br
V-17: R1 = CHF2, R2 = 2-Br
V-18: R1 = CF3, R2 = 2-Br
V-19: R1 = Me, R2 = 4-i-Pr
V-20: R1 = CHF2, R2 = 4-i-Pr
V-21: R1 = CF3, R2 = 4-i-Pr
V-22: R1 = Me, R2 = 3, 4-Cl2
V-23: R1 = CHF2, R2 = 3, 4-Cl2
V-24: R1 = CF3, R2 = 3, 4-Cl2
V-25: R1 = Me, R2 = 4-Et
V-26: R1 = CHF2, R2 = 4-Et
V-27: R1 = CF3, R2 = 4-Et
V-28: R1 = Me, R2 = 4-n-Pr
V-29: R1 = CHF2, R2 = 4-n-Pr
V-30: R1 = CF3, R2 = 4-n-Pr
V-31: R1 = Me, R2 = 4-t-Bu
V-32: R1 = CHF2, R2 = 4-t-Bu
V-33: R1 = CF3, R2 = 4-t-Bu
Scheme 1. Synthesis of compound V. Reagents and conditions: (a) potassium carbonate, hydrazine
hydrate, tetrahydrofuran, 0–25 °C; (b) sodium hydride, ethyl acetate/ethyl ether, -5–25 °C; (c) sul-
furic acid, ethanol, 75 °C; (d) sulfuric acid, H2O, 100 °C .
2.2. Docking Analysis
Molecular docking was used to predict the binding modes and molecular interactions
of compound V by MOE (Version 2020.09). The binding energy was predicted based on
the structure and configurations of the compounds, as summarized in Table 1. The bind-
ing energies of almost all target compounds were less than that of picloram, which indi-
cated that most of target compound V exhibited a higher affinity for AFB5. In particular,
the binding energy of compound V-7 (−8.59 kJ mol−1) was the lowest.
Table 1. Predicted binding energy of all target compounds, picloram, and halauxifen-methyl.
Compd.
Score (kJ mol−1)
Compd.
Score (kJ mol−1)
V-1
−8.07
V-19
−7.77
V-2
−8.33
V-20
−7.58
V-3
−8.15
V-21
−7.04
V-4
−8.02
V-22
−7.88
V-5
−8.00
V-23
−7.90
V-6
−8.09
V-24
−7.81
V-7
−8.59
V-25
−8.07
V-8
−8.29
V-26
−7.97
V-9
−8.30
V-27
−7.90
Scheme 1.
Synthesis of compound
V
. Reagents and conditions: (
a
) potassium carbonate, hy-
drazine hydrate, tetrahydrofuran, 0–25
◦
C; (
b
) sodium hydride, ethyl acetate/ethyl ether,
−
5–25
◦
C;
(c) sulfuric acid, ethanol, 75 ◦C; (d) sulfuric acid, H2O, 100 ◦C.
2.2. Docking Analysis
Molecular docking was used to predict the binding modes and molecular interactions
of compound
V
by MOE (Version 2020.09). The binding energy was predicted based on the
structure and configurations of the compounds, as summarized in Table 1. The binding
energies of almost all target compounds were less than that of picloram, which indicated
that most of target compound
V
exhibited a higher affinity for AFB5. In particular, the
binding energy of compound V-7 (−8.59 kJ mol−1) was the lowest.
As shown in Figure 4, compound
V-7
, whose binding energy was
−
8.33 kJ mol
−1
,
exhibited hydrogen bonding with five amino acid residues: Arg449, Arg482, Arg123,
Phe127, and Asp126, whereas picloram with a binding energy of
−
6.53 kJ mol
−1
exhibited
hydrogen bonding with three amino acid residues: Arg449, Val485, and Leu450, which
probably explained the difference of the binding energies between two molecules at a certain
level. In particular, the nitrogen atom at position 2 of the pyrazolyl ring in compound V-7
forms a hydrogen bond with residues Arg123 and Arg 482, which demonstrates that the
proposed design improved the modification of molecules.
Molecules 2023,28, 1431 5 of 20
Table 1. Predicted binding energy of all target compounds, picloram, and halauxifen-methyl.
Compd. Score (kJ mol−1) Compd. Score (kJ mol−1)
V-1 −8.07 V-19 −7.77
V-2 −8.33 V-20 −7.58
V-3 −8.15 V-21 −7.04
V-4 −8.02 V-22 −7.88
V-5 −8.00 V-23 −7.90
V-6 −8.09 V-24 −7.81
V-7 −8.59 V-25 −8.07
V-8 −8.29 V-26 −7.97
V-9 −8.30 V-27 −7.90
V-10 −7.54 V-28 −7.36
V-11 −7.80 V-29 −7.53
V-12 −7.80 V-30 −7.62
V-13 −8.23 V-31 −7.44
V-14 −8.29 V-32 −6.81
V-15 −8.03 V-33 −7.28
V-16 −8.09 Picloram −6.53
V-17 −8.05 Halauxifen-methyl −7.25
V-18 −8.11
Use London dG and GBVI/WSA dG as rescoring functions.
Molecules 2023, 28, x FOR PEER REVIEW 5 of 21
V-10
−7.54
V-28
−7.36
V-11
−7.80
V-29
−7.53
V-12
−7.80
V-30
−7.62
V-13
−8.23
V-31
−7.44
V-14
−8.29
V-32
−6.81
V-15
−8.03
V-33
−7.28
V-16
−8.09
Picloram
−6.53
V-17
−8.05
Halauxifen-methyl
−7.25
V-18
−8.11
Use London dG and GBVI/WSA dG as rescoring functions.
As shown in Figure 4, compound V-7, whose binding energy was −8.33 kJ mol−1, ex-
hibited hydrogen bonding with five amino acid residues: Arg449, Arg482, Arg123,
Phe127, and Asp126, whereas picloram with a binding energy of −6.53 kJ mol−1 exhibited
hydrogen bonding with three amino acid residues: Arg449, Val485, and Leu450, which
probably explained the difference of the binding energies between two molecules at a cer-
tain level. In particular, the nitrogen atom at position 2 of the pyrazolyl ring in compound
V-7 forms a hydrogen bond with residues Arg123 and Arg 482, which demonstrates that
the proposed design improved the modification of molecules.
(a)
(b)
(c)
(d)
Figure 4. AFB5 exhibits different auxin-binding affinities to compounds. Three-dimensional struc-
tures of AFB5 exhibit an almost identical fold with regard to secondary structure arrangements. (a)
Docking arrangements of compound V-7 to AFB5. (b) Docking arrangements of picloram to AFB5.
White sticks represent hydrogen atoms, red sticks represent oxygen atoms, blue sticks represent
Figure 4.
AFB5 exhibits different auxin-binding affinities to compounds. Three-dimensional struc-
tures of AFB5 exhibit an almost identical fold with regard to secondary structure arrangements.
(a) Docking
arrangements of compound
V-7
to AFB5. (
b
) Docking arrangements of picloram to AFB5.
White sticks represent hydrogen atoms, red sticks represent oxygen atoms, blue sticks represent
nitrogen atoms and dark green sticks represent chlorine atoms. (
c
) Two-dimensional interaction of
compound V-7 to AFB5. (d) Two-dimensional interaction of picloram to AFB5.
Molecules 2023,28, 1431 6 of 20
As reported by Jeffrey B. Epp et al., halauxinfen-methyl is metabolized into halauxifen
in plants [
13
]. In comparison with the binding to AFB5, as shown in Figure 5b, compound
V-7
and halauxifen overlapped well in the same position, and compound
V-7
not only
exhibited more hydrogen bonding, but also formed hydrophobic interactions between
the benzene ring and the nearby phenylalanine because of the lengthening effect of the
pyrazolyl ring.
Molecules 2023, 28, x FOR PEER REVIEW 6 of 21
nitrogen atoms and dark green sticks represent chlorine atoms. (c) Two-dimensional interaction of
compound V-7 to AFB5. (d) Two-dimensional interaction of picloram to AFB5.
As reported by Jeffrey B. Epp et al., halauxinfen-methyl is metabolized into halauxi-
fen in plants [13]. In comparison with the binding to AFB5, as shown in Figure 5b, com-
pound V-7 and halauxifen overlapped well in the same position, and compound V-7 not
only exhibited more hydrogen bonding, but also formed hydrophobic interactions be-
tween the benzene ring and the nearby phenylalanine because of the lengthening effect of
the pyrazolyl ring.
(a)
(b)
(c)
Figure 5. (a) Docking arrangements of halauxifen to AFB5. (b) Superposition diagram of compound
V-7 and halauxifen docking arrangements to AFB5. White sticks represent hydrogen atoms, red
sticks represent oxygen atoms, blue sticks represent nitrogen atoms and dark green sticks represent
chlorine atoms. (c) Two-dimensional interaction of halauxifen to AFB5.
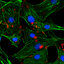
2.3. A. thaliana Root Growth Assays to Quantify Compounds Activity
A. thaliana is frequently used as a model plant to explore the preliminary effects of
novel compounds on plants [28], including phenotype and physiological indexes, and it
can also be used to study the mechanism of plant responses to chemicals at the protein
and gene levels. In this study, all target compounds were tested against A. thaliana root
growth in at least five concentrations for preliminary biological activity and the calcula-
tion of IC50 values. As summarized in Table 2, some compounds exhibited better inhibition
than the picloram and halauxifen-methyl commercial herbicides. In particular, com-
pounds V-2 and V-7 exhibited approximately 30- and 50-fold lower IC50 values than the
Figure 5.
(
a
) Docking arrangements of halauxifen to AFB5. (
b
) Superposition diagram of compound
V-7
and halauxifen docking arrangements to AFB5. White sticks represent hydrogen atoms, red sticks
represent oxygen atoms, blue sticks represent nitrogen atoms and dark green sticks represent chlorine
atoms. (c) Two-dimensional interaction of halauxifen to AFB5.
2.3. A. thaliana Root Growth Assays to Quantify Compounds Activity
A. thaliana is frequently used as a model plant to explore the preliminary effects of
novel compounds on plants [
28
], including phenotype and physiological indexes, and it
can also be used to study the mechanism of plant responses to chemicals at the protein and
gene levels. In this study, all target compounds were tested against A. thaliana root growth
in at least five concentrations for preliminary biological activity and the calculation of IC
50
values. As summarized in Table 2, some compounds exhibited better inhibition than the
picloram and halauxifen-methyl commercial herbicides. In particular, compounds
V-2
and
V-7
exhibited approximately 30- and 50-fold lower IC
50
values than the commercial herbi-
cide halauxifen-methyl, respectively, indicating that the proposed strategy for modifying
picloram is probably successful in inhibiting A. thaliana root growth.
Molecules 2023,28, 1431 7 of 20
Table 2. IC50 values of all compounds against A. thaliana root growth inhibition.
Compd. IC50 (µmol/L) Compd. IC50 (µmol/L)
V-1 2.1172 V-19 20.3215
V-2 0.0688 V-20 36.5507
V-3 1.1394 V-21 43.2911
V-4 1.4501 V-22 6.7232
V-5 2.1794 V-23 6.4282
V-6 1.8578 V-24 7.0823
V-7 0.0389 V-25 1.3623
V-8 1.4044 V-26 4.1381
V-9 0.8444 V-27 2.2558
V-10 25.2618 V-28 33.9772
V-11 7.1501 V-29 44.886
V-12 7.6716 V-30 23.0342
V-13 0.8872 V-31 27.6596
V-14 1.2416 V-32 73.1934
V-15 1.6909 V-33 57.9380
V-16 0.1850 Picloram 3.7240
V-17 0.6585 Halauxifen-methyl 1.7764
V-18 0.7218
All target compounds were tested using at least five concentrations with triplicates to determine IC50 values.
Combined with the score and pIC
50
, as shown in Figure 6, the scatter plots of score and
pIC50 are generally fitted to a line, indicating that the docking model was well predicted.
Molecules 2023, 28, x FOR PEER REVIEW 7 of 21
commercial herbicide halauxifen-methyl, respectively, indicating that the proposed strat-
egy for modifying picloram is probably successful in inhibiting A. thaliana root growth.
Table 2. IC50 values of all compounds against A. thaliana root growth inhibition.
Compd.
IC50 (μmol/L)
Compd.
IC50 (μmol/L)
V-1
2.1172
V-19
20.3215
V-2
0.0688
V-20
36.5507
V-3
1.1394
V-21
43.2911
V-4
1.4501
V-22
6.7232
V-5
2.1794
V-23
6.4282
V-6
1.8578
V-24
7.0823
V-7
0.0389
V-25
1.3623
V-8
1.4044
V-26
4.1381
V-9
0.8444
V-27
2.2558
V-10
25.2618
V-28
33.9772
V-11
7.1501
V-29
44.886
V-12
7.6716
V-30
23.0342
V-13
0.8872
V-31
27.6596
V-14
1.2416
V-32
73.1934
V-15
1.6909
V-33
57.9380
V-16
0.1850
Picloram
3.7240
V-17
0.6585
Halauxifen-methyl
1.7764
V-18
0.7218
All target compounds were tested using at least five concentrations with triplicates to determine
IC50 values.
Combined with the score and pIC50, as shown in Figure 6, the scatter plots of score
and pIC50 are generally fitted to a line, indicating that the docking model was well pre-
dicted.
Figure 6. Scatter plots of the score (the predicted binding energy) versus pIC50.
2.4. Three-Dimensional Quantitative Structure–Activity Relationship (3D-QSAR)
Combined with the determination of IC50 values against A. thaliana root growth inhi-
bition and compound structure, a 3D-QSAR model was constructed using the CoMFA
strategy. The model was generated with all possible combinations of steric and electro-
static fields for CoMFA [29]. To build the model, the cross-validated partial least squares
(PLS) method was used, which provides a leave-one-out cross-validated correlation
Figure 6. Scatter plots of the score (the predicted binding energy) versus pIC50.
2.4. Three-Dimensional Quantitative Structure–Activity Relationship (3D-QSAR)
Combined with the determination of IC
50
values against A. thaliana root growth
inhibition and compound structure, a 3D-QSAR model was constructed using the CoMFA
strategy. The model was generated with all possible combinations of steric and electrostatic
fields for CoMFA [
29
]. To build the model, the cross-validated partial least squares (PLS)
method was used, which provides a leave-one-out cross-validated correlation coefficient q
2
and determination coefficient r
2
. Based on the basic requirements of the criteria proposed
by the Golbraikh and Tropsha conditions, the model satisfying q
2
> 0.5 and r
2
> 0.6 is
considered to be acceptable and predictive.
As listed in Table 3, the CoMFA model satisfied the q
2
> 0.5 and r
2
> 0.6 criteria.
This model exhibited a q
2
of 0.679; r
2
of 0.848; standard error of estimate (SEE) of 0.337;
Fisher test value (F) of 44.660; and an optimum number of components (ONC) of 4. The
contributions of steric and electrostatic fields were 62.7 and 36.3%, respectively.
Molecules 2023,28, 1431 8 of 20
Table 3. Statistical result of the CoMFA model.
Model q2r2SEE FONC Field Contribution (%)
S E
CoMFA 0.679 0.848 0.337 44.660 4 62.7 36.3
Leave-one-out cross-validated correlation coefficient (q
2
), determination coefficient (r
2
), error of estimate (SEE),
Fisher’s test value (F), optimum number of components (ONC), steric (S), and electrostatic (E).
2.4.1. Scatter Plots
As shown in Figure 7, the scatter plots of the actual and predicted inhibitory activities
of the training set compounds fit well to a line, indicating that the constructed CoMFA
model exhibits predictive capability.
Molecules 2023, 28, x FOR PEER REVIEW 8 of 21
coefficient q2 and determination coefficient r2. Based on the basic requirements of the cri-
teria proposed by the Golbraikh and Tropsha conditions, the model satisfying q2 > 0.5 and
r2 > 0.6 is considered to be acceptable and predictive.
As listed in Table 3, the CoMFA model satisfied the q2 > 0.5 and r2 > 0.6 criteria. This
model exhibited a q2 of 0.679; r2 of 0.848; standard error of estimate (SEE) of 0.337; Fisher
test value (F) of 44.660; and an optimum number of components (ONC) of 4. The contri-
butions of steric and electrostatic fields were 62.7 and 36.3%, respectively.
Table 3. Statistical result of the CoMFA model.
Model
q2
r2
SEE
F
ONC
Field Contribution (%)
S
E
CoMFA
0.679
0.848
0.337
44.660
4
62.7
36.3
Leave-one-out cross-validated correlation coefficient (q2), determination coefficient (r2), error of es-
timate (SEE), Fisher’s test value (F), optimum number of components (ONC), steric (S), and electro-
static (E).
2.4.1. Scatter Plots
As shown in Figure 7, the scatter plots of the actual and predicted inhibitory activities
of the training set compounds fit well to a line, indicating that the constructed CoMFA
model exhibits predictive capability.
Figure 7. Scatter plots of the actual versus predicted pIC50 for the CoMFA model.
2.4.2. Contour Map Analysis
The CoMFA contour map was created using the StDev*Coeff mapping option, which
allows visualization of each field effect. Compound V-7, which had the lowest IC50 value,
was superimposed on the contour map. The contributions to the favorable and unfavora-
ble regions in each field were 80 and 20%, respectively.
In the steric contour map shown in Figure 8a, the green contour block indicates that
the presence of bulky steric groups in this area would be favorable for the biological ac-
tivity of a compound, whereas the yellow block indicates that the bulky groups would be
unfavorable. The green contour blocks appear in four parts around compound V-7, while
the largest green contour block appears at the fourth position of the benzene ring con-
nected to the pyrazole ring, which indicates the reason for the different activities between
compounds V-7 and V-1. Their substituents are chlorine atoms and methyl groups, re-
spectively, and chlorine atom groups exhibit larger occupation than methyl groups. In
addition, it can be observed in this figure that a small yellow contour block appears next
to the largest green contour block, indicating that the bulky groups in this area are unfa-
vorable. This could explain why compound V-7 was more active than compound V-32,
because in comparison with chlorine atoms, the tert-butyl group occupies a larger space,
resulting in its invasion of the area of the yellow contour block.
-2
-1
0
1
2
-2 -1 012
Predicated pIC50
Actual pIC50
Figure 7. Scatter plots of the actual versus predicted pIC50 for the CoMFA model.
2.4.2. Contour Map Analysis
The CoMFA contour map was created using the StDev*Coeff mapping option, which
allows visualization of each field effect. Compound
V-7
, which had the lowest IC
50
value,
was superimposed on the contour map. The contributions to the favorable and unfavorable
regions in each field were 80 and 20%, respectively.
In the steric contour map shown in Figure 8a, the green contour block indicates that the
presence of bulky steric groups in this area would be favorable for the biological activity of a
compound, whereas the yellow block indicates that the bulky groups would be unfavorable.
The green contour blocks appear in four parts around compound
V-7
, while the largest
green contour block appears at the fourth position of the benzene ring connected to the
pyrazole ring, which indicates the reason for the different activities between compounds
V-7
and
V-1
. Their substituents are chlorine atoms and methyl groups, respectively, and
chlorine atom groups exhibit larger occupation than methyl groups. In addition, it can be
observed in this figure that a small yellow contour block appears next to the largest green
contour block, indicating that the bulky groups in this area are unfavorable. This could
explain why compound
V-7
was more active than compound
V-32
, because in comparison
with chlorine atoms, the tert-butyl group occupies a larger space, resulting in its invasion
of the area of the yellow contour block.
As shown in Figure 8b, the blue contours indicate that the electropositive groups are
favorable for increasing the activity of a compound, whereas the red contours indicate
that the electropositive groups are unfavorable for the activity of a compound. On the
side of the carbonyl group, a large red contour block appears, indicating the importance
of the carbonyl group at position 2 of the pyridine. The blue contour block next to the
trifluoromethyl group at position 3 of the pyrazolyl ring explains why compound
V-7
exhibits higher activity owing to the presence of electronegative substituents in the blue
contour region.
Molecules 2023,28, 1431 9 of 20
Molecules 2023, 28, x FOR PEER REVIEW 9 of 21
As shown in Figure 8b, the blue contours indicate that the electropositive groups are
favorable for increasing the activity of a compound, whereas the red contours indicate
that the electropositive groups are unfavorable for the activity of a compound. On the side
of the carbonyl group, a large red contour block appears, indicating the importance of the
carbonyl group at position 2 of the pyridine. The blue contour block next to the trifluoro-
methyl group at position 3 of the pyrazolyl ring explains why compound V-7 exhibits
higher activity owing to the presence of electronegative substituents in the blue contour
region.
(a)
(b)
Figure 8. Contour maps of CoMFA analysis with the most active compound V-7. In each field, fa-
vored and disfavored areas are fixed at 80 and 20% contribution levels, respectively; (a) steric field:
green contours represent the favored regions whereas yellow contours represent the disfavored re-
gions; (b) electrostatic field: blue contours represent the favored regions whereas red contours rep-
resent the disfavored regions.
2.5. Greenhouse Activity Assay
Based on the design strategy and docking analysis, the herbicidal activities of the new
compounds were tested in a greenhouse against six common weeds, including three gra-
mineous weeds: Setaria glauca (SG), Digitaria sanguinalis (DS), and Echinochloa crusgalli
(EC); three broadleaf weeds: Chenopodium album (CA), Abutilon theophrasti (AT), and Am-
aranthus retroflexus (AR); and two commercial herbicides, picloram and halauxifen-me-
thyl, were selected as the control groups. As summarized in Table 4 and shown in Figure
9, most of the compounds exhibited post-emergence inhibitory activities against the
weeds mentioned above. Generally, the inhibitory activities of almost all compounds on
broadleaf weeds are stronger than those on gramineous weeds, which is similar to the
inhibitory activities of picloram and halauxifen-methyl. This also indicates that the target
compounds may have similar mechanisms of action as commercial herbicides. Further-
more, compounds V-1–V-18 with halogen-substituted phenyl exhibited better herbicidal
activities than compounds V-19–V-33 with alkyl-substituted phenyl. Notably, compound
V-8 at a dosage of 300 g ha−1 exhibits 100, 100, and 95% post-emergence injury values
against CA, AT, and AR, respectively, and 100, 40, and 100% post-emergence injury values
against CA, AT, and AR, respectively. This illustrates that the herbicidal activity of com-
pound V-8 was better than that of picloram. Furthermore, there was a slight variance be-
tween the herbicidal activities of compounds V and inhibition of A. thaliana root growth,
possibly because of the AFB5 difference in A. thaliana and the tested weeds.
Figure 8.
Contour maps of CoMFA analysis with the most active compound
V-7
. In each field,
favored and disfavored areas are fixed at 80 and 20% contribution levels, respectively; (
a
) steric
field: green contours represent the favored regions whereas yellow contours represent the disfavored
regions; (
b
) electrostatic field: blue contours represent the favored regions whereas red contours
represent the disfavored regions.
2.5. Greenhouse Activity Assay
Based on the design strategy and docking analysis, the herbicidal activities of the new
compounds were tested in a greenhouse against six common weeds, including three grami-
neous weeds: Setaria glauca (SG), Digitaria sanguinalis (DS), and Echinochloa crusgalli (EC);
three broadleaf weeds: Chenopodium album (CA), Abutilon theophrasti (AT), and Amaranthus
retroflexus (AR); and two commercial herbicides, picloram and halauxifen-methyl, were
selected as the control groups. As summarized in Table 4and shown in Figure 9, most of the
compounds exhibited post-emergence inhibitory activities against the weeds mentioned
above. Generally, the inhibitory activities of almost all compounds on broadleaf weeds
are stronger than those on gramineous weeds, which is similar to the inhibitory activities
of picloram and halauxifen-methyl. This also indicates that the target compounds may
have similar mechanisms of action as commercial herbicides. Furthermore, compounds
V-1
–
V-18
with halogen-substituted phenyl exhibited better herbicidal activities than com-
pounds
V-19
–
V-33
with alkyl-substituted phenyl. Notably, compound
V-8
at a dosage of
300 gha−1
exhibits 100, 100, and 95% post-emergence injury values against CA, AT, and
AR, respectively, and 100, 40, and 100% pre-emergence injury values against CA, AT, and
AR, respectively. This illustrates that the herbicidal activity of compound
V-8
was better
than that of picloram. Furthermore, there was a slight variance between the herbicidal
activities of compounds
V
and inhibition of A. thaliana root growth, possibly because of the
AFB5 difference in A. thaliana and the tested weeds.
Based on these results, the herbicidal activity of compound
V-8
was tested on broadleaf
weeds at dosages of 300, 150, and 75 gha
−1
. As summarized in Table 5, the inhibition of
compound
V-8
against the weeds gradually decreased with decreasing dosages in the
pre-emergence test; however, the decreasing trend was small. In the post-emergence test,
the inhibition of compound
V-8
remained high at a dosage of 150 gha
−1
compared with
that at a dosage of 300 gha
−1
. These results indicated that compound
V-8
exhibited a better
herbicidal effect than picloram.
Molecules 2023,28, 1431 10 of 20
Table 4. Herbicidal activities of all compounds.
Compd. Dosage
(gha−1)
Post-Emergence Pre-Emergence
SG DS EC CA AT AR SG DS EC CA AT AR
V-1 300 50 10 0 80 0 90 70 0 0 0 0 20
V-2 300 0 0 0 75 10 90 15 0 0 95 0 95
V-3 300 10 0 0 100 65 100 60 0 0 0 0 60
V-4 300 20 0 0 95 90 95 30 40 0 100 0 95
V-5 300 25 0 0 90 10 80 30 0 0 60 0 60
V-6 300 60 20 15 100 0 100 35 10 0 30 0 75
V-7 300 0 0 0 100 0 100 10 0 0 90 20 85
V-8 300 40 0 20 100 100 95 20 0 0 100 40 100
V-9 300 70 0 50 90 0 80 20 0 0 10 0 20
V-10 300 85 50 0 100 10 70 80 60 0 60 10 20
V-11 300 70 0 0 60 0 40 60 40 0 90 20 30
V-12 300 0 0 0 40 0 0 30 0 0 0 20 0
V-13 300 0 0 0 80 60 85 0 0 0 20 0 60
V-14 300 40 0 0 100 60 100 0 0 0 90 0 25
V-15 300 35 0 0 75 0 95 0 0 0 0 0 0
V-16 300 35 0 0 50 55 95 0 0 0 100 0 80
V-17 300 40 0 0 85 20 90 10 0 0 0 0 90
V-18 300 50 0 0 85 50 90 0 0 0 0 0 0
V-19 300 50 0 0 10 25 0 0 0 0 0 0 0
V-20 300 30 0 0 10 20 0 0 0 0 0 0 30
V-21 300 55 0 0 10 15 0 0 0 0 0 0 0
V-22 300 20 0 0 10 10 0 0 0 0 0 0 0
V-23 300 40 0 0 10 0 0 0 0 0 0 0 0
V-24 300 3000000000000
V-25 300 40 0 0 10 0 15 0 0 0 30 0 0
V-26 300 50 0 0 40 0 0 10 0 0 30 0 0
V-27 300 6000000000000
V-28 300 70 0 0 10 50 0 0 0 0 0 0 0
V-29 300 60 0 0 0 10 15 0 0 0 0 0 0
V-30 300 20 0 0 0 10 0 0 0 0 0 0 0
V-31 300 20 0 0 0 10 0 0 0 0 0 0 0
V-32 300 70 0 0 10 10 0 0 0 0 0 0 0
V-33 300 3000000000000
Picloram 300 45 65 0 90 80 90 25 0 0 90 30 75
Halauxifen-
methyl 15 30 40 60 90 80 98 0 0 0 0 20 40
All compounds with picloram and halauxifen-methyl as controls were evaluated by percent visual injury value
against SG, DS, EC, CA, AT, AR under pre-emergence and post-emergence conditions.
Table 5. Herbicidal activities of compound V-8.
Dosage (gha−1)Post-Emergence Pre-Emergence
CA AT AR CA AT AR
300 / / / 100.0 59.0 98.0
150 93.9 35.7 100 98.0 35.9 95.0
75 / / / 95.0 7.0 81.2
Evaluation of fresh weight inhibition in the aboveground against CA, AT, AR under pre-emergence and post-
emergence conditions.
To further investigate the crop selectivity of compound
V-8
, three common
crops—including
corn, wheat, and sorghum under pre-emergence and post-emergence
conditions—were treated. As summarized in Table 6, compound
V-8
was safe for wheat
while slightly damaging corn and sorghum under pre-emergence conditions; however, it
was completely safe for the three crops under post-emergence conditions.
Molecules 2023,28, 1431 11 of 20
Molecules 2023, 28, x FOR PEER REVIEW 11 of 21
Figure 9. Accumulated visual injury of compound V and picloram under post-emergence treatment.
Based on these results, the herbicidal activity of compound V-8 was tested on broad-
leaf weeds at dosages of 300, 150, and 75 g ha−1. As summarized in Table 5, the inhibition
of compound V-8 against the weeds gradually decreased with decreasing dosages in the
pre-emergence test; however, the decreasing trend was small. In the post-emergence test,
the inhibition of compound V-8 remained high at a dosage of 150 g ha−1 compared with
that at a dosage of 300 g ha−1. These results indicated that compound V-8 exhibited a better
herbicidal effect than picloram.
Table 5. Herbicidal activities of compound V-8.
Dosage (g ha−1)
Post-Emergence
Pre-Emergence
CA
AT
AR
CA
AT
AR
300
/
/
/
100.0
59.0
98.0
150
93.9
35.7
100
98.0
35.9
95.0
75
/
/
/
95.0
7.0
81.2
Evaluation of fresh weight inhibition in the aboveground against CA, AT, AR under pre-emergence
(pre) and post-emergence (post) conditions.
To further investigate the crop selectivity of compound V-8, three common crops—
including corn, wheat, and sorghum under pre-emergence (pre) and post-emergence
(post) conditions—were treated. As summarized in Table 6, compound V-8 was safe for
wheat while slightly damaging corn and sorghum under pre-emergence conditions; how-
ever, it was completely safe for the three crops under post-emergence conditions.
Table 6. Crop selectivity of compound V-8.
Dosage (g ha−1)
Post-Emergence
Pre-Emergence
Corn
Wheat
Sorghum
Corn
Wheat
Sorghum
300
0
0
0
13.4
0
14.0
Fresh weight inhibition in the aboveground parts of corn, wheat, and sorghum under pre-emer-
gence (pre) and post-emergence (post) conditions.
Figure 9.
Accumulated visual injury of compound
V
and picloram under post-emergence treatment.
Table 6. Crop selectivity of compound V-8.
Dosage (gha−1)Post-Emergence Pre-Emergence
Corn Wheat Sorghum Corn Wheat Sorghum
300 0 0 0 13.4 0 14.0
Fresh weight inhibition in the aboveground parts of corn, wheat, and sorghum under pre-emergence (pre) and
post-emergence (post) conditions.
In summary, these results demonstrated that the target compounds are potential
candidates for herbicide discovery research, and they need to be studied further.
3. Materials and Methods
3.1. Chemicals and Instruments
All the reagents and solvents were purchased from commercial suppliers (Beijing
InnoChem Science & Technology Co., Ltd., Beijing, China and Sinopharm Chemical Reagent
Co., Ltd., Beijing, China). All reactions were monitored using thin-layer chromatography
(TLC) run on silica gel glass plates (Qingdao Broadchem Industrial, Qingdao, China).
1
H
and
13
C NMR spectroscopy were recorded using a Bruker AM-500 spectrometer with
temperature control at 21–23
◦
C, using DMSO-d6 or CDCl
3
as the solvent and tetramethyl
silane (TMS) as the internal reference. In the spectra, the chemical shifts (
δ
) were given in
parts per million (ppm). High-resolution mass spectra (HRMS) were determined with an
Agilent 6540 QTOF instrument.
3.2. Synthesis
3.2.1. General Synthetic Procedure of Intermediate II
In a 500 mL, three-necked, round-bottom flask, compound
I
(100 mmol) and anhydrous
potassium carbonate (200 mmol) were added to tetrahydrofuran (250 mL). Subsequently,
80% hydrazine hydrate (300 mmol) was slowly added to the reaction mixture while stirring
at 0
◦
C. After the addition, the reaction mixture was heated to 75
◦
C in an oil bath for 6 h
of reflux. Once the reaction was complete, the mixture was cooled to room temperature
and filtered. The obtained solid was washed with water to obtain intermediate
II
as an
off-white solid (yield 73%):
1
H NMR (300 MHz, DMSO-d6)
δ
9.63 (s, 1H), 7.18 (s, 2H), 4.53
(s, 2H).
Molecules 2023,28, 1431 12 of 20
3.2.2. General Synthetic Procedure of Intermediate III
Two different procedures were used to synthesize 1,3-diketones in this study.
(1) When
R
1
was a methyl group, ethyl acetate was used as the reactant and solvent. In a
100 mL
round-bottom flask, 60% sodium hydride (28.8 mmol) was slowly added to 30 mL of ethyl
acetate under stirring at
−
5
◦
C. Thereafter, a mixture of methyl ketones (28.8 mmol) and
10 mL of ethyl acetate was added slowly to the reaction solution. After addition, the
entire system was warmed to room temperature and stirred for 6 h. The reaction was
quenched with aqueous hydrochloric acid (1 N, 30 mL), acidified to a pH range of 1–2, and
subsequently extracted using ethyl acetate (3
×
15 mL). The combined organic phases were
dried over anhydrous sodium sulfate and concentrated under a vacuum. The residue was
purified via flash column chromatography (n-hexane) to obtain intermediate
III
(yields
65.8–78.4%). (2) When R
1
was a difluoromethyl or trifluoromethyl, the procedure was as
follows: in a 100 mL round-bottom flask, 60% sodium hydride (43.2 mmol) was slowly
added to 30 mL of ethyl ether with stirring at
−
5
◦
C. Thereafter, a mixture of methyl
ketones (28.8 mmol) and ester (34.56 mmol) was slowly added to the reaction solution.
After addition, the entire system was warmed to room temperature and stirred overnight.
The reaction was quenched with aqueous hydrochloric acid (1 N, 30 mL), acidified to a pH
range of 1–2, and extracted using ethyl acetate (3
×
15 mL). The combined organic phases
were dried over anhydrous sodium sulfate and concentrated under vacuum. The residue
was purified via flash column chromatography (n-hexane/ethyl acetate = 10:1) to afford
intermediate III (yields 87.6–92.2%).
3.2.3. General Synthetic Procedure of Intermediate IV
In a 50 mL round-bottom flask, intermediate
III
(5 mmol) was added to a solution of
intermediate
II
(5 mmol) in 20 mL of ethanol at room temperature. Thereafter, concentrated
sulfuric acid was slowly added to the stirred solution, and the reaction mixture was heated
to 75
◦
C in an oil bath and kept at reflux for 2 h. The reaction was cooled to room temper-
ature, quenched with a saturated sodium carbonate solution, and extracted using ethyl
acetate (3
×
15 mL). The combined organic phases were dried over anhydrous sodium
sulfate and concentrated under vacuum. The residue was purified via flash column chro-
matography (n-hexane/ethyl acetate = 6:1) to afford intermediate IV (yields 80.3–91.7%).
3.2.4. General Synthetic Procedure of Compound V
Intermediate
IV
(1.067 mmol) was dissolved in 80% aqueous sulfuric acid (10 mL) in a
25 mL round-bottom flask. Thereafter, the reaction solution was heated to 100
◦
C in an oil
bath and kept at reflux for 2 h. The reaction mixture was cooled to room temperature and
quenched with water. The white solid was collected through filtration and dried to achieve
the target compound V(yields 90.1–99.0%).
Compound
V-1
4-Amino-3,5-dichloro-6-(3-methyl-5-(p-tolyl)-1H-pyrazol-1-yl)picolinic
acid. White solid.
1
H NMR (500 MHz, DMSO-d6)
δ
13.90 (s, 1H), 7.21 (s, 2H), 7.14 (d,
J= 8.1 Hz
, 2H), 7.08 (d, J= 8.1 Hz, 2H), 6.47 (s, 1H), 2.25 (s, 3H), 2.25 (s, 3H).
13
C NMR
(126 MHz, DMSO-d6)
δ
165.94, 150.50, 149.53, 147.69, 147.12, 145.13, 138.41, 129.79, 127.35,
127.13, 113.10, 112.26, 106.17, 21.19, 13.79. HRMS calcd. for C
17
H
14
Cl
2
N
4
O
2
([M-H]
−
),
375.0416; found, 375.0413.
Compound
V-2
4-Amino-3,5-dichloro-6-(3-(difluoromethyl)-5-(p-tolyl)-1H-pyrazol-1-
yl)picolinic acid. White solid.
1
H NMR (300 MHz, DMSO-d6)
δ
14.03 (s, 1H), 7.35 (s, 2H),
7.19 (d, J= 8.6 Hz, 2H), 7.16 (d, J= 8.6 Hz, 2H), 7.10 (t, J= 54.35 Hz, 1H), 6.99 (s, 1H), 2.27 (s,
3H).
13
C NMR (126 MHz, DMSO-d6)
δ
165.70, 163.80, 161.84, 150.85, 148.00, 147.78, 147.54,
147.30, 146.49, 145.07, 130.24, 130.17, 125.32, 116.60, 116.43, 113.53, 113.01, 112.72, 111.67,
109.82, 104.16. HRMS calcd. for C17H12Cl2F2N4O2([M-H]−), 411.0227; found, 411.0239.
Compound
V-3
4-Amino-3,5-dichloro-6-(5-(p-tolyl)-3-(trifluoromethyl)-1H-pyrazol-
1-yl)picolinic acid. White solid.
1
H NMR (500 MHz, DMSO-d6)
δ
14.04 (s, 1H), 7.40 (s,
2H), 7.25 (s, 1H), 7.21 (d, J= 8.9 Hz, 2H), 7.18 (d, J= 8.9 Hz, 2H), 2.28 (s, 3H).
13
C NMR
(126 MHz, DMSO-d6)
δ
165.73, 150.83, 147.51, 146.63, 146.38, 143.22, 142.92, 142.62, 142.32,
Molecules 2023,28, 1431 13 of 20
139.77, 130.04, 127.85, 125.32, 124.88, 122.74, 120.60, 118.46, 113.17, 112.73, 104.51, 40.30,
21.22. HRMS calcd. for C17H11Cl2F3N4O2([M-H]−), 429.0133; found, 429.0143.
Compound
V-4
4-Amino-3,5-dichloro-6-(5-(4-fluorophenyl)-3-methyl-1H-pyrazol-1-
yl)picolinic acid. Yellow solid.
1
H NMR (500 MHz, DMSO-d6)
δ
13.83 (s, 1H), 7.25 (s,
2H), 7.24–7.22 (m, 2H), 7.22–7.18 (m, 2H), 6.52 (s, 1H), 2.26 (s, 3H).
13
C NMR (126 MHz,
DMSO-d6)
δ
165.88, 163.36, 161.41, 150.61, 149.65, 147.36, 147.15, 144.06, 129.71, 129.64,
126.50, 116.37, 116.19, 112.90, 112.32, 106.68, 13.76. HRMS calcd. for C
16
H
11
Cl
2
FN
4
O
2
([M-H]−), 379.0165; found, 379.0159.
Compound
V-5
4-Amino-3,5-dichloro-6-(3-(difluoromethyl)-5-(4-fluorophenyl)-1H-
pyrazol-1-yl)picolinic acid. Yellow solid.
1
H NMR (500 MHz, DMSO-d6)
δ
13.98 (s, 1H),
7.40 (s, 2H), 7.35–7.33 (m, 2H), 7.28–7.24 (m, 2H), 7.13 (t, J= 54.3 Hz, 1H), 7.07 (s, 1H).
13
C
NMR (126 MHz, DMSO-d6)
δ
165.70, 163.80, 161.84, 150.85, 148.00, 147.78, 147.54, 147.30,
146.49, 145.07, 130.24, 130.17, 125.32, 116.60, 116.43, 113.53, 113.01, 112.72, 111.67, 109.82,
104.16. HRMS calcd. for C16H9Cl2F3N4O2([M-H]−), 414.9976; found, 414.9986.
Compound
V-6
4-Amino-3,5-dichloro-6-(5-(4-fluorophenyl)-3-(trifluoromethyl)-1H-
pyrazol-1-yl)picolinic acid. Yellow solid.
1
H NMR (500 MHz, DMSO-d6)
δ
14.07 (s, 1H), 7.46
(s, 2H), 7.38–7.36 (m, 2H), 7.35 (s, 1H), 7.31–7.27 (m, 2H).
13
C NMR (126 MHz, DMSO-d6)
δ
165.61, 164.01, 162.03, 150.97, 147.30, 146.07, 145.56, 143.29, 142.99, 142.69, 142.39, 130.44,
130.37, 124.81, 124.71, 124.68, 124.68, 122.67, 120.53, 118.39, 116.71, 116.54, 113.34, 112.67,
105.06. HRMS calcd. for C16H8Cl2F4N4O2([M-H]−), 432.9882; found, 432.9888.
Compound
V-7
4-Amino-3,5-dichloro-6-(5-(4-chlorophenyl)-3-methyl-1H-pyrazol-1-
yl)picolinic acid. White solid.
1
H NMR (500 MHz, DMSO-d6)
δ
13.50 (s, 1H), 7.43 (d,
J= 8.55 Hz
, 2H), 7.28 (s, 2H), 7.21 (d, J= 8.55 Hz, 2H), 6.57 (s, 1H), 2.27 (s, 3H).
13
C NMR
(126 MHz, DMSO-d6)
δ
165.85, 150.66, 149.78, 147.28, 147.11, 143.85, 133.63, 129.34, 129.16,
128.79, 112.80, 112.39, 106.96, 13.76. HRMS calcd. for C
16
H
10
Cl
3
N
4
O
2
([M-H]
−
), 394.9869;
found, 395.0023.
Compound
V-8
4-Amino-3,5-dichloro-6-(5-(4-chlorophenyl)-3-(difluoromethyl)-1H-
pyrazol-1-yl)picolinic acid. White solid.
1
H NMR (300 MHz, DMSO-d6)
δ
13.95 (s, 1H),
7.48 (d, J= 8.4 Hz, 2H), 7.40 (s, 2H),7.30 (d, J= 8.4 Hz, 2H), 7.14 (t, J= 54.3 Hz, 1H), 7.11 (s,
1H).
13
C NMR (126 MHz, DMSO-d6)
δ
165.65, 150.88, 148.08, 147.85, 147.62, 147.31, 146.41,
144.87, 134.49, 129.61, 129.54, 127.61, 113.47, 113.07, 111.62, 109.77, 104.43. HRMS calcd. for
C16H8Cl3F2N4O2([M-H]−), 430.9681; found, 430.9688.
Compound
V-9
4-Amino-3,5-dichloro-6-(5-(4-chlorophenyl)-3-(trifluoromethyl)-1H-
pyrazol-1-yl)picolinic acid. White solid.
1
H NMR (500 MHz, DMSO-d6)
δ
14.06 (s, 1H), 7.51
(d, J= 8.5 Hz, 2H), 7.47 (s, 2H), 7.38 (s, 1H), 7.33 (d, J= 8.5 Hz, 2H).
13
C NMR (126 MHz,
DMSO-d6)
δ
165.61, 151.00, 147.34, 146.00, 145.36, 143.36, 143.06, 142.76, 142.46, 134.90,
129.75, 129.62, 126.99, 124.77, 122.63, 120.49, 118.35, 113.39, 112.58, 105.32. HRMS calcd. for
C16H7Cl3F3N4O2([M-H]−), 448.9587; found, 448.9592.
Compound
V-10
4-Amino-3,5-dichloro-6-(5-(3-chlorophenyl)-3-methyl-1H-pyrazol-1-
yl)picolinic acid. Yellow solid.
1
H NMR (500 MHz, Chloroform-d)
δ
7.29 (s, 1H), 7.28 (d,
J= 7.75 Hz
, 1H), 7.23 (dd, J= 8.0 Hz, 7.75 Hz, 1H), 6.99 (d, J= 7.75 Hz, 1H), 6.40 (s, 1H),
5.62 (s, 2H), 2.41 (s, 3H).
13
C NMR (126 MHz, DMSO-d6)
δ
165.85, 150.64, 149.79, 147.23,
143.48, 133.82, 131.85, 131.13, 128.77, 127.35, 125.91, 112.85, 112.34, 107.27, 13.77. HRMS
calcd. for C16H10Cl3N4O2([M-H]−), 394.9869; found, 394.9873.
Compound
V-11
4-Amino-3,5-dichloro-6-(5-(3-chlorophenyl)-3-(difluoromethyl)-1H-
pyrazol-1-yl)picolinic acid. Yellow solid.
1
H NMR (500 MHz, DMSO-d6)
δ
13.96 (s, 1H),
7.46–7.45 (m, 1H), 7.45 (s, 1H), 7.43–7.40 (m, 1H), 7.39 (s, 2H), 7.18 (s, 1H), 7.16–7.13 (m,
1H), 7.14 (t, J= 54.2 Hz, 1H).
13
C NMR (126 MHz, DMSO-d6)
δ
165.64, 150.87, 148.10,
147.87, 147.64, 147.38, 146.39, 144.50, 134.01, 131.28, 130.68, 129.58, 127.88, 126.30, 113.45,
113.04, 112.71, 111.60, 109.75, 104.81. HRMS calcd. for C
16
H
8
Cl
3
F
2
N
4
O
2
([M-H]
−
), 430.9681;
found, 430.9691.
Compound V-12 4-Amino-3,5-dichloro-6-(5-(3-chlorophenyl)-3-(trifluoromethyl)-1H-
pyrazol-1-yl)picolinic acid. Yellow solid.
1
H NMR (500 MHz, DMSO-d6)
δ
14.01 (s, 1H),
7.52–7.39 (m, 6H), 7.15 (d, J= 7.7 Hz, 1H).
13
C NMR (126 MHz, DMSO-d6)
δ
165.58, 151.00,
Molecules 2023,28, 1431 14 of 20
147.29, 145.97, 144.98, 143.37, 143.07, 142.77, 142.47, 134.10, 131.36, 130.01, 129.95, 128.09,
126.36, 125.69, 124.74, 124.60, 122.60, 120.46, 118.33, 113.36, 112.67, 105.70. HRMS calcd. for
C16H7Cl3F3N4O2([M-H]−), 448.9587; found, 448.9592.
Compound
V-13
4-Amino-6-(5-(4-bromophenyl)-3-methyl-1H-pyrazol-1-yl)-3,5-
dichloropicolinic acid. Yellow solid.
1
H NMR (500 MHz, DMSO-d6)
δ
13.85 (s, 1H),
7.56 (d, J= 8.2 Hz, 2H), 7.23 (s, 2H), 7.14 (d, J= 8.3 Hz, 2H), 6.57 (s, 1H), 2.27 (s, 3H).
13
C
NMR (126 MHz, DMSO-d6)
δ
165.90, 150.62, 149.77, 147.25, 143.89, 132.26, 129.41, 129.15,
122.28, 112.67, 112.30, 106.93, 13.76. HRMS calcd. for C
16
H
10
BrCl
2
N
4
O
2
([M-H]
−
), 440.9344;
found, 440.9348.
Compound
V-14
4-Amino-6-(5-(4-bromophenyl)-3-(difluoromethyl)-1H-pyrazol-1-yl)-
3,5-dichloropicolinic acid. Yellow solid.
1
H NMR (500 MHz, DMSO-d6)
δ
13.92 (s, 1H), 7.62
(d, J= 8.5 Hz, 1H), 7.38 (s, 2H), 7.23 (d, J= 8.5 Hz, 1H), 7.13 (t, J= 54.2 Hz, 1H), 7.11 (s,
1H).
13
C NMR (126 MHz, DMSO-d6)
δ
165.61, 150.89, 148.10, 147.88, 147.65, 147.26, 146.41,
144.93, 132.45, 129.83, 127.97, 123.21, 113.47, 113.10, 112.64, 111.61, 109.76, 104.43. HRMS
calcd. for C16H8BrCl2F2N4O2([M-H]−), 476.9155; found, 476.9159.
Compound
V-15
4-Amino-6-(5-(4-bromophenyl)-3-(trifluoromethyl)-1H-pyrazol-1-yl)-
3,5-dichloropicolinic acid. Yellow solid.
1
H NMR (500 MHz, DMSO-d6)
δ
13.92 (s, 1H), 7.57
(d, J= 8.5 Hz, 2H), 7.39 (s, 2H), 7.31 (s, 1H), 7.18 (d, J= 8.5 Hz, 2H).
13
C NMR (126 MHz,
DMSO-d6)
δ
165.58, 151.01, 147.27, 145.99, 145.42, 143.38, 143.08, 142.77, 142.48, 132.54,
129.95, 127.33, 124.76, 123.64, 122.62, 120.48, 118.34, 113.42, 112.58, 105.32. HRMS calcd. for
C16H7BrCl2F3N4O2([M-H]−), 494.9061; found, 494.9066.
Compound
V-16
4-Amino-6-(5-(2-bromophenyl)-3-methyl-1H-pyrazol-1-yl)-3,5-
dichloropicolinic acid. Yellow solid.
1
H NMR (500 MHz, DMSO-d6)
δ
13.71 (s, 1H),
7.66 (dd, J= 7.9, 1.1 Hz, 1H), 7.31 (td, J= 7.5, 1.2 Hz, 1H), 7.26 (td, J= 7.7, 1.8 Hz, 1H),
7.16 (s, 2H), 7.15 (dd, J= 7.7, 1.8 Hz, 1H), 6.48 (s, 1H), 2.30 (s, 3H).
13
C NMR (126 MHz,
DMSO-d6)
δ
165.79, 150.43, 149.03, 146.67, 146.64, 143.25, 133.51, 132.02, 131.26, 130.98,
127.84, 123.02, 112.09, 111.70, 109.10, 13.83. HRMS calcd. for C
16
H
10
BrCl
2
N
4
O
2
([M-H]
−
),
440.9344; found, 440.9351.
Compound
V-17
4-Amino-6-(5-(2-bromophenyl)-3-(difluoromethyl)-1H-pyrazol-1-yl)-
3,5-dichloropicolinic acid. Yellow solid.
1
H NMR (500 MHz, DMSO-d6)
δ
13.82 (s, 1H), 7.71
(dd, J= 7.7, 1.1 Hz, 1H), 7.38–7.27 (m, 4H), 7.22 (dd, J= 7.3, 2.0 Hz, 1H), 7.17 (t, J= 54.2 Hz,
1H), 6.98 (s, 1H).
13
C NMR (126 MHz, DMSO-d6)
δ
165.61, 150.66, 147.38, 147.15, 146.92,
146.85, 145.86, 144.28, 133.60, 132.27, 131.73, 129.98, 128.01, 123.25, 113.51, 112.45, 112.13,
111.66, 109.81, 106.57, 40.30. HRMS calcd. for C
16
H
8
BrCl
2
F
2
N
4
O
2
([M-H]
−
), 476.9155;
found, 476.9158.
Compound
V-18
4-Amino-6-(5-(2-bromophenyl)-3-(trifluoromethyl)-1H-pyrazol-1-yl)-
3,5-dichloropicolinic acid. Yellow solid.
1
H NMR (500 MHz, DMSO-d6)
δ
13.91 (s, 1H), 7.73
(dd, J= 7.6, 1.5 Hz, 1H), 7.37 (m, 4H), 7.26(dd, J= 7.3, 2.0 Hz, 1H), 7.25 (s, 1H).
13
C NMR
(126 MHz, DMSO-d6)
δ
165.53, 150.77, 146.84, 145.45, 144.81, 142.67, 142.37, 142.07, 141.77,
133.62, 132.37, 132.08, 129.32, 128.07, 124.80, 123.36, 122.66, 120.52, 118.38, 112.78, 112.16,
107.41. HRMS calcd. for C16H7BrCl2F3N4O2([M-H]−), 494.9061; found, 494.9064.
Compound
V-19
4-Amino-3,5-dichloro-6-(5-(4-isopropylphenyl)-3-methyl-1H-pyrazol-
1-yl)picolinic acid. Yellow solid.
1
H NMR (500 MHz, DMSO-d6)
δ
13.78 (s, 1H), 7.23 (s,
2H), 7.21 (d, J= 8.4 Hz, 2H), 7.13 (d, J= 8.3 Hz, 2H), 6.48 (s, 1H), 2.84 (hept, J= 6.9 Hz,
1H), 2.25 (s, 3H), 1.16 (d, J= 6.9 Hz, 6H).
13
C NMR (126 MHz, DMSO-d6)
δ
165.92, 150.52,
149.50, 149.12, 147.78, 147.12, 145.03, 127.36, 127.19, 113.23, 112.31, 106.24, 33.52, 24.05, 13.78.
HRMS calcd. for C19H17Cl2N4O2([M-H]−), 403.0729; found, 403.0734.
Compound
V-20
4-Amino-3,5-dichloro-6-(3-(difluoromethyl)-5-(4-isopropylphenyl)-
1H-pyrazol-1-yl)picolinic acid. White solid.
1
H NMR (500 MHz, DMSO-d6)
δ
13.95 (s, 1H),
7.36 (s, 2H), 7.27 (d, J= 8.3 Hz, 2H), 7.22 (d, J= 8.3 Hz, 2H), 7.11 (t, J= 54.3 Hz, 1H), 7.01 (s,
1H), 2.86 (hept, J= 6.9 Hz, 1H), 1.17 (d, J= 6.9 Hz, 6H).
13
C NMR (126 MHz, DMSO-d6)
δ
166.42, 150.45, 149.91, 147.80, 147.57, 147.34, 146.81, 145.97, 127.74, 127.38, 126.33, 113.63,
112.33, 112.16, 111.78, 109.93, 103.57, 33.56, 24.01. HRMS calcd. for C
19
H
15
Cl
2
F
2
N
4
O
2
([M-H]−), 439.0540; found, 439.0550.
Molecules 2023,28, 1431 15 of 20
Compound
V-21
4-Amino-3,5-dichloro-6-(5-(4-isopropylphenyl)-3-(trifluoromethyl)-
1H-pyrazol-1-yl)picolinic acid. Yellow solid.
1
H NMR (500 MHz, DMSO-d6)
δ
13.98 (s, 1H),
7.43 (s, 2H), 7.34–7.21 (m, 5H), 2.87 (hept, J= 6.9 Hz, 1H), 1.17 (d, J= 6.9 Hz, 5H).
13
C NMR
(126 MHz, DMSO-d6)
δ
165.68, 150.89, 150.40, 147.31, 146.54, 146.48, 143.24, 142.94, 142.64,
142.34, 127.88, 127.47, 125.64, 124.88, 122.74, 120.60, 118.46, 113.28, 112.90, 104.60, 33.59,
23.97. HRMS calcd. for C19H14Cl2F3N4O2([M-H]−), 457.0446; found, 457.0461.
Compound
V-22
4-Amino-3,5-dichloro-6-(5-(3,4-dichlorophenyl)-3-methyl-1H-pyrazol-
1-yl)picolinic acid. White solid.
1
H NMR (500 MHz, DMSO-d6)
δ
13.89 (s, 1H), 7.62 (d,
J= 8.4 Hz
, 1H), 7.55 (d, J= 2.0 Hz, 1H), 7.27 (s, 2H), 7.05 (dd, J= 8.4, 2.1 Hz, 1H), 6.68 (s,
1H), 2.27 (s, 3H).
13
C NMR (126 MHz, DMSO-d6)
δ
165.86, 150.71, 149.88, 147.37, 147.00,
142.49, 131.98, 131.58, 131.50, 130.40, 129.41, 127.30, 112.65, 112.39, 107.61, 13.75. HRMS
calcd. for C16H9Cl4N4O2([M-H]−), 430.9450; found, 430.9460.
Compound
V-23
4-Amino-3,5-dichloro-6-(5-(3,4-dichlorophenyl)-3-(difluoromethyl)-
1H-pyrazol-1-yl)picolinic acid. White solid.
1
H NMR (500 MHz, DMSO-d6)
δ
14.03 (s, 1H),
7.69 (d, J= 2.1 Hz, 1H), 7.67 (d, J= 8.4 Hz, 1H), 7.42 (s, 2H), 7.23 (s, 1H), 7.14 (t, J= 54.2 Hz,
1H), 7.12 (dd, J= 8.4, 2.1 Hz, 1H).
13
C NMR (126 MHz, DMSO-d6)
δ
165.66, 150.96, 148.15,
147.92, 147.69, 147.41, 146.16, 143.52, 132.49, 132.21, 131.65, 130.02, 129.20, 127.62, 113.40,
113.12, 112.55, 111.54, 109.69, 105.15, 40.30. HRMS calcd. for C
16
H
7
Cl
4
F
2
N
4
O
2
([M-H]
−
),
466.9262; found, 466.9273.
Compound V-24 4-Amino-3,5-dichloro-6-(5-(3,4-dichlorophenyl)-3-(trifluoromethyl)-
1H-pyrazol-1-yl)picolinic acid. White solid.
1
H NMR (500 MHz, DMSO-d6)
δ
14.11 (s,
1H), 7.74 (d, J= 2.1 Hz, 1H), 7.69 (d, J= 8.4 Hz, 1H), 7.48 (s, 1H), 7.45 (s, 2H), 7.14 (dd,
J= 8.4
, 2.1 Hz, 1H).
13
C NMR (126 MHz, DMSO-d6)
δ
165.69, 151.02, 147.80, 145.74, 144.00,
143.38, 143.08, 142.78, 142.48, 132.90, 132.30, 131.73, 130.24, 128.55, 127.70, 124.69, 122.55,
120.41, 118.27, 113.33, 112.39, 105.99. HRMS calcd. for C
16
H
6
Cl
4
F
3
N
4
O
2
([M-H]
−
), 484.9176;
found, 484.9181.
Compound
V-25
4-Amino-3,5-dichloro-6-(5-(4-ethylphenyl)-3-methyl-1H-pyrazol-1-
yl)picolinic acid. White solid.
1
H NMR (500 MHz, DMSO-d6)
δ
13.22 (s, 1H),7.21 (s, 2H),
7.17 (d, J= 8.2 Hz, 2H), 7.12 (d, J= 8.2 Hz, 2H), 6.48 (s, 1H), 2.56 (q, J= 7.6 Hz, 2H), 1.14
(t, J= 7.6 Hz, 3H).
13
C NMR (126 MHz, DMSO-d6)
δ
165.94, 150.51, 149.52, 147.72, 147.12,
145.09, 144.55, 128.60, 127.39, 127.34, 113.15, 112.27, 106.21, 28.22, 15.58, 13.78. HRMS calcd.
for C18H15Cl2N4O2([M-H]−), 389.0572; found, 389.0584.
Compound
V-26
4-Amino-3,5-dichloro-6-(3-(difluoromethyl)-5-(4-ethylphenyl)-1H-
pyrazol-1-yl)picolinic acid. White solid.
1
H NMR (500 MHz, DMSO-d6)
δ
14.05 (s, 1H), 7.37
(s, 2H), 7.23 (d, J= 8.4 Hz, 2H), 7.20 (d, J= 8.4 Hz, 2H), 7.11 (t, J= 54.2 Hz, 1H), 7.00 (s, 1H),
2.59 (q, J= 7.6 Hz, 2H), 1.15 (t, J= 7.6 Hz, 3H).
13
C NMR (126 MHz, DMSO)
δ
166.37, 150.46,
147.82, 147.59, 147.36, 146.76, 146.06, 145.36, 128.76, 127.77, 126.23, 113.63, 112.37, 112.18,
111.78, 109.93, 103.54, 40.28, 28.24, 15.51. HRMS calcd. for C
18
H
13
Cl
2
F
2
N
4
O
2
([M-H]
−
),
425.0384; found, 425.0394.
Compound
V-27
4-Amino-3,5-dichloro-6-(5-(4-ethylphenyl)-3-(trifluoromethyl)-1H-
pyrazol-1-yl)picolinic acid. White solid.
1
H NMR (500 MHz, DMSO-d6)
δ
13.95 (s, 1H),
7.41 (s, 2H), 7.26 (s, 1H), 7.24 (d, J= 8.3 Hz, 2H), 7.22 (d, J= 8.3 Hz, 2H), 2.59 (q, J= 7.6 Hz,
2H), 1.15 (t, J= 7.6 Hz, 3H).
13
C NMR (126 MHz, DMSO-d6)
δ
165.67, 150.86, 147.27, 146.61,
146.40, 145.88, 143.23, 142.94, 142.64, 142.34, 128.86, 127.89, 125.50, 124.86, 122.72, 120.58,
118.44, 113.25, 112.85, 104.55, 28.23, 15.48. HRMS calcd. for C
18
H
12
Cl
2
F
3
N
4
O
2
([M-H]
−
),
443.0289; found, 443.0306.
Compound
V-28
4-Amino-3,5-dichloro-6-(3-methyl-5-(4-propylphenyl)-1H-pyrazol-
1-yl)picolinic acid. White solid.
1
H NMR (500 MHz, DMSO-d6)
δ
14.14 (s, 1H), 7.21 (s,
2H), 7.15 (d, J= 8.3 Hz, 2H), 7.11 (d, J= 8.3 Hz, 2H), 6.48 (s, 1H), 2.51 (t, J= 7.6 Hz, 2H),
2.26 (s, 3H), 1.61–1.49 (m, 2H), 0.85 (t, J= 7.3 Hz, 3H).
13
C NMR (126 MHz, DMSO-d6)
δ
165.94, 150.51, 149.52, 147.71, 147.12, 145.10, 142.99, 129.15, 127.37, 127.31, 113.11, 112.24,
106.20, 37.29, 24.17, 14.11, 13.79. HRMS calcd. for C
19
H
17
Cl
2
N
4
O
2
([M-H]
−
), 403.0729;
found, 403.0742.
Molecules 2023,28, 1431 16 of 20
Compound
V-29
4-Amino-3,5-dichloro-6-(3-(difluoromethyl)-5-(4-propylphenyl)-1H-
pyrazol-1-yl)picolinic acid. White solid.
1
H NMR (500 MHz, DMSO-d6)
δ
14.18 (s, 1H), 7.25
(s, 2H), 7.19 (t, J= 1.7 Hz, 4H), 7.10 (t, J= 54.2 Hz, 1H), 6.99 (s, 1H), 2.51 (t, J= 7.6 Hz, 2H),
1.60–1.50 (m, 2H), 0.85 (t, J= 7.3 Hz, 3H).
13
C NMR (126 MHz, DMSO-d6)
δ
165.75, 150.74,
147.93, 147.71, 147.47, 147.31, 146.83, 146.11, 143.88, 129.34, 127.69, 126.20, 113.59, 112.88,
111.74, 109.89, 103.65, 37.28, 24.12, 14.10. HRMS calcd. for C
19
H
15
Cl
2
F
2
N
4
O
2
([M-H]
−
),
439.0540; found, 439.0554.
Compound
V-30
4-Amino-3,5-dichloro-6-(5-(4-propylphenyl)-3-(trifluoromethyl)-1H-
pyrazol-1-yl)picolinic acid. Yellow solid.
1
H NMR (500 MHz, DMSO-d6)
δ
14.13 (s, 1H), 7.42
(s, 2H), 7.26 (s, 1H), 7.22 (s, 4H), 2.57–2.50 (m, 2H), 1.56 (m, 2H), 0.86 (t, J= 7.3 Hz, 3H).
13
C
NMR (126 MHz, DMSO-d6)
δ
165.66, 150.87, 147.28, 146.62, 146.41, 144.31, 143.23, 142.93,
142.64, 142.33, 129.41, 127.82, 125.56, 124.87, 122.74, 120.60, 118.46, 113.22, 112.83, 104.55,
37.28, 24.09, 14.07. HRMS calcd. for C
19
H
14
Cl
2
F
3
N
4
O
2
([M-H]
−
), 457.0446;
found, 457.0459.
Compound
V-31
4-Amino-6-(5-(4-(tert-butyl)phenyl)-3-methyl-1H-pyrazol-1-yl)-3,5-
dichloropicolinic acid. White solid.
1
H NMR (500 MHz, DMSO-d6)
δ
13.89 (s, 1H), 7.36 (d,
J= 8.4 Hz, 2H), 7.24 (s, 2H), 7.15 (d, J= 8.4 Hz, 2H), 6.49 (s, 1H), 2.25 (s, 3H), 1.24 (s, 9H).
13C NMR (126 MHz, DMSO)
δ
165.94, 151.39, 150.54, 149.50, 147.82, 147.13, 144.88, 127.04,
126.08, 113.27, 112.35, 106.27, 40.14, 34.86, 31.39, 13.78. HRMS calcd. for C
20
H
19
Cl
2
N
4
O
2
([M-H]−), 417.0885; found, 417.0903.
Compound
V-32
4-Amino-6-(5-(4-(tert-butyl)phenyl)-3-(difluoromethyl)-1H-pyrazol-
1-yl)-3,5-dichloropicolinic acid. White solid.
1
H NMR (500 MHz, DMSO-d6)
δ
13.92 (s, 1H),
7.77 (s, 2H), 7.42 (d, J= 8.5 Hz, 2H), 7.23 (d, J= 8.5 Hz, 2H), 7.11 (t, J= 54.2 Hz, 1H), 7.05 (s,
1H), 1.25 (s, 9H).
13
C NMR (126 MHz, DMSO-d6)
δ
166.91, 152.13, 150.16, 147.67, 147.44,
147.21, 146.77, 145.76, 127.43, 126.25, 126.04, 113.66, 111.81, 111.78, 111.45, 109.97, 34.93,
31.35. HRMS calcd. for C20H17Cl2F2N4O2([M-H]−), 453.0697; found, 453.0713.
Compound
V-33
4-Amino-6-(5-(4-(tert-butyl)phenyl)-3-(trifluoromethyl)-1H-pyrazol-
1-yl)-3,5-dichloropicolinic acid. White solid.
1
H NMR (500 MHz, DMSO-d6)
δ
13.99 (s,
1H), 7.43 (d, J= 8.5 Hz, 2H), 7.32 (s, 2H), 7.27 (d, J= 8.6 Hz, 3H), 1.25 (s, 9H).
13
C NMR
(126 MHz, DMSO-d6)
δ
166.52, 152.57, 150.42, 146.40, 146.30, 143.03, 142.73, 142.43, 142.13,
127.57, 126.35, 125.51, 125.37, 124.93, 122.79, 120.65, 118.52, 112.35, 111.71, 104.46, 34.97,
31.32. HRMS calcd. for C20H16Cl2F3N4O2([M-H]−), 471.0602; found, 471.0620.
3.3. Homology Modeling of AFB5
The crystal structure of A. Thaliana AFB5 protein has not yet been verified; therefore,
we conducted homology modeling based on TIR1 as the template protein from Protein Data
Bank (PDB) coded 3C6O: chain B. The sequence of AFB5 obtained from TAIR (https://www.
arabidopsis.org/, accessed on 10 October 2022) has a similarity of 50.98% with that of TIR1;
therefore, it is feasible to construct the AFB5 protein structure upon TIR1 through homology
modeling. The AFB5 protein was evaluated using SAVES6.0 (
SAVESv6.0—Structure
Val-
idation Server (ucla.edu)) and SWISS MODEL (
https://swissmodel.expasy.org/assess/
,
accessed on 15 October 2022) after optimizing the individual residues of the modeled
protein using MOE. As shown in Figure 10, the Ramachandran plot showed 96.59% of all
backbone dihedral angles in favored areas; 95.61% of the residues had an averaged 3D-1D
score of
≥
0.2, and the number of non-bond interactions formed between pairs of different
atomic types on the side chain in the 3.5 nm range overall quality factor was 88.59 (
≥
0.2).
The QMEAN value was
−
2.62, whereas the GMQE value was 0.80, which verified that the
AFB5 protein modeled using MOE is of good quality.
3.4. Molecular Docking
Two-dimensional structures of picolinic acid derivatives were generated using Chem-
Draw Professional 16.0; their configurations were minimized and protonated, and charge
was applied to the 3D structures using MOE2020. Thereafter, the protein was protonated,
charge was added, and 4.5 Å of water molecules near the pocket was eliminated before
molecular docking. The docking pocket and key residues were reported by Calderón
Molecules 2023,28, 1431 17 of 20
Villalobos, which were put forward to the dock. The side chains near the pocket were set as
free rotation, and refinement was set as an induced fit. London dG and GBVI/WSA dG
were used as the rescoring functions. A total of 500 conformations of each compound were
generated to predict their best possible binding pose and output 20 tops-core optimum con-
figurations, which could be browsed on MOE, balancing score, and key binding residues to
choose one conformation and output as the final conformation.
Molecules 2023, 28, x FOR PEER REVIEW 17 of 21
147.44, 147.21, 146.77, 145.76, 127.43, 126.25, 126.04, 113.66, 111.81, 111.78, 111.45, 109.97,
34.93, 31.35. HRMS calcd. for C20H17Cl2F2N4O2 ([M-H]−), 453.0697; found, 453.0713.
Compound V-33 4-Amino-6-(5-(4-(tert-butyl)phenyl)-3-(trifluoromethyl)-1H-pyra-
zol-1-yl)-3,5-dichloropicolinic acid. White solid. 1H NMR (500 MHz, DMSO-d6) δ 13.99 (s,
1H), 7.43 (d, J = 8.5 Hz, 2H), 7.32 (s, 2H), 7.27 (d, J = 8.6 Hz, 3H), 1.25 (s, 9H). 13C NMR (126
MHz, DMSO-d6) δ 166.52, 152.57, 150.42, 146.40, 146.30, 143.03, 142.73, 142.43, 142.13,
127.57, 126.35, 125.51, 125.37, 124.93, 122.79, 120.65, 118.52, 112.35, 111.71, 104.46, 34.97,
31.32. HRMS calcd. for C20H16Cl2F3N4O2 ([M-H]−), 471.0602; found, 471.0620.
3.3. Homology Modeling of AFB5
The crystal structure of A. Thaliana AFB5 protein has not yet been verified; therefore,
we conducted homology modeling based on TIR1 as the template protein from Protein
Data Bank (PDB) coded 3C6O: chain B. The sequence of AFB5 obtained from TAIR
(https://www.arabidopsis.org/, accessed on 10 October 2022) has a similarity of 50.98%
with that of TIR1; therefore, it is feasible to construct the AFB5 protein structure upon
TIR1 through homology modeling. The AFB5 protein was evaluated using SAVES6.0
(SAVESv6.0—Structure Validation Server (ucla.edu)) and SWISS MODEL (https://swiss-
model.expasy.org/assess/, accessed on 15 October 2022) after optimizing the individual
residues of the modeled protein using MOE. As shown in Figure 10, the Ramachandran
plot showed 96.59% of all backbone dihedral angles in favored areas; 95.61% of the resi-
dues had an averaged 3D-1D score of ≥0.2, and the number of non-bond interactions
formed between pairs of different atomic types on the side chain in the 3.5nm range over-
all quality factor was 88.59 (≥0.2). The QMEAN value was −2.62, whereas the GMQE value
was 0.80, which verified that the AFB5 protein modeled using MOE is of good quality.
Figure 10. Ramachandran plots of the AFB5 structural model.
3.4. Molecular Docking
Two-dimensional structures of picolinic acid derivatives were generated using
ChemDraw Professional 16.0; their configurations were minimized and protonated, and
charge was applied to the 3D structures using MOE2020. Thereafter, the protein was pro-
tonated, charge was added, and 4.5 Å of water molecules near the pocket was eliminated
before molecular docking. The docking pocket and key residues were reported by Calde-
rón Villalobos, which were put forward to the dock. The side chains near the pocket were
set as free rotation, and refinement was set as an induced fit. London dG and GBVI/WSA
dG were used as the rescoring functions. A total of 500 conformations of each compound
were generated to predict their best possible binding pose and output 20 tops-core
Figure 10. Ramachandran plots of the AFB5 structural model.
3.5. Biological Assay
3.5.1. Root Growth Assays to Quantify Compounds Activity
The designed and synthesized compounds possess skeleton 2-picolinic acid and could
exhibit the same inhibition on Arabidopsis thaliana root growth as auxin. Therefore, they
were assayed for their influence on Arabidopsis thaliana root growth to explore preliminary
bioactivity and evaluate their IC
50
values. Arabidopsis thaliana seeds were surface-sterilized
and spotted onto 1/2
×
Murashige and Skoog medium containing 0.7% agar, 3% sucrose,
and compounds at the indicated concentrations in petri dishes. Subsequently, the dishes
were transferred to the dark at 4
◦
C for 48 h. After that, the dishes were placed vertically
into the incubator for 7 d at 22
◦
C for 16 h:8 h (day/night). The root lengths of 7-day-old
seedlings were measured using IMAGEJ after the images were acquired.
The inhibition rates of root growth were determined as follows:
P=
L0−L
L0
×100%,
where P denotes the inhibition rate, and L and L
0
are the average length of the A. thaliana
root in the presence of compounds and in untreated controls, respectively.
Determination of the IC
50
values was performed using an Internet tool: MLA—”Quest
Graph
™
IC
50
Calculator.” AAT Bioquest, Inc., 4 November 2022, https://www.aatbio.com/
tools/ic50-calculator.
3.5.2. 3D-QSAR
Thirty-seven compounds, including thirty-three target compounds and four Yang
compounds, were used for the 3D-QSAR study. Their structures and inhibitory activities
are listed in Table 2. To develop the CoMFA model, all the compounds mentioned were
used as a training set based on the structural and bioactive diversity using the DIVERSITY
function of SYBYL-X 2.0 (Tripos, Inc., St. Louis, MO, USA). The compound structures in
the test set sufficiently represent the diversity of the entire dataset. A training set was used
to construct the model.
Molecules 2023,28, 1431 18 of 20
All compounds in the training set were drawn in ChemDraw and superimposed
and aligned on the maximum common substructure of 6-(5-aryl-substituted-pyrazolyl)-
2-picolinic acid derivatives using the alignment function in SYBYL-X 2.0. Based on the
hypothesis that the common alignment core contributes equally to the bioactivity of the
compounds, the conformational angles for the maximum common substructures of the
most active compound
V-7
as the template for the alignment were copied and applied to
the remaining compounds in the whole dataset. Figure 8shows the most active compound
V-7
as the template, and the structural alignment of the training set for the 3D-QSAR study.
CoMFA was performed using the Lennard-Jones potential for the steric field and the
Coulombic potential for the electrostatic field in SYBYL-X 2.0. The aligned compounds
were placed in a 3D cubic lattice with 2.0 Å grid spacing in the x, y, and z directions, and
these potentials were determined for each compound. A sp3 carbon probe with a van der
Waals radius of 1.52 Å and a point charge of +1.0 was used at each lattice point of the
grid box for the calculation of the steric and electrostatic fields. To avoid overpower due
to the large steric and electrostatic energy values, the default energy cutoff value was set
at 30 kcal/mol. The attenuation coefficient was set to a default value of 0.3. Regression
analysis was performed using the cross-validated PLS method. To develop the final model
by performing a PLS analysis, the first run was conducted through the cross-validation to
obtain the ONC, and thereafter, the final run yielded the non-cross-validated r2value.
3.5.3. Greenhouse Herbicidal Activity Assay
Herbicidal activities were evaluated at the Collaborative Innovation Center for Green
Pesticides, Zhejiang A&F University (Hangzhou, China), and the soil for herbicidal activity
tests was collected from a local planted field. All compounds were dissolved in 100% DMSO
and thereafter diluted with 0.1% Tween-80 solution to obtain the appropriate concentrations
before use. The pre- and post-emergence herbicidal activities of the compounds were
evaluated against six weeds, including three dicotyledonous weeds: SG, DS, and EC, and
three gramineous weeds: CA, AT, and AR. In a greenhouse, the weed seeds were planted
in a plastic pot with a diameter of 8.0 cm at the mouth of the pot, covered with 0.2–0.5 cm
of soil after seeding, and the bottom was watered and cultured in a plant culture room at a
temperature of 13
±
8
◦
C. For the post-emergence herbicidal activity assays, the spray was
applied at the 2–4 leaf stage of the weeds. For the pre-emergence herbicidal activity assay,
the spray was applied 24 h after weed seeding. After the weed treatments, the weeds were
transferred to a greenhouse for cultivation with standard management. Weed growth and
toxic symptoms were observed regularly after treatment, and weed inhibitory activities
were visually evaluated (two duplicates per experiment) 20 days after treatment to obtain
the percent visual injury value. In the further herbicidal activity assay of compound V-8,
fresh weight inhibition (three triplicates per experiment) in the aboveground was applied
rather than visual observation. All sprays were performed using a Biotest Spray Tower
(3WPSH-500E, Nanjing Institute of Agricultural Mechanization, Ministry of Agriculture
and Rural Affairs).
3.5.4. Crop Selectivity
In the greenhouse, three representative crops (corn, wheat, and sorghum) were used
to evaluate the crop selectivity of the compounds using the procedure described above.
4. Conclusions
In this study, 33 4-amino-3,5-dicholor-6-(5-aryl-substituted-1-pyrazolyl)-2-picolinic
acid compounds were designed and synthesized via a four-step synthetic route with
good yields based on the splicing of the active fragments, wherein the phenyl-substituted
pyrazolyl replaced the chlorine atom at position 6 of picloram. The docking analysis
demonstrated that some compounds might be bioactive owing to their tighter affinity to
AFB5 of A. thaliana. The primary bioassay for inhibiting A. thaliana root growth demon-
strated that the IC
50
value of compound
V-7
was 45-fold less than that of the commercial
Molecules 2023,28, 1431 19 of 20
picloram herbicide. Based on this, a 3D-QSAR model was constructed and fitted well to the
relationship between the bioactivity and structure, which could be used to design new lead
compounds. The herbicidal activity tested in greenhouses demonstrated that compound
V-8
exhibited better post-emergence herbicidal activity against broadleaf weeds at a dosage
of 300 gha
−1
than picloram. A crop selectivity test indicated that compound
V-8
exhibited
excellent crop safety against corn, wheat, and sorghum at a dosage of 300 gha
−1
under
post-emergence conditions. Nevertheless, the actual herbicidal activity of the compounds
was different from the primary bioactive results, while the docking analysis was fit to A.
thaliana root growth assays, which may be due to the biological differences between weeds
and A. thaliana; therefore, the results were acceptable. These results demonstrated that the
replacement of the chlorine atom at position 6 of picloram by phenyl-substituted pyrazolyl
is favorable for improving the herbicidal activity of skeleton picloram, and compound
V-8
might be a potential lead structure for the discovery of novel synthetic auxin herbicides.
Furthermore, these results provide new perspectives and insights for the future design of
compounds with similar structures. Further studies on structural optimization and active
mechanisms are currently in progress in our laboratory.
Supplementary Materials:
The following supporting information can be downloaded at: https://
www.mdpi.com/article/10.3390/molecules28031431/s1.
1
H and
13
C NMR spectra of compounds
V-1–V-33.
Author Contributions:
Conceptualization, Y.-M.C. and S.-Z.L.; methodology, T.F. and Z.-Y.X.; soft-
ware L.Z., Q.L. and T.F.; bioassay, T.F., Q.L. and W.W.; validation, H.-T.L. and R.-C.S. All authors have
read and agreed to the published version of the manuscript.
Funding:
This research was funded by the National Key R&D Program, grant number 2022YFD1700403.
Institutional Review Board Statement: Not applicable.
Informed Consent Statement: Not applicable.
Data Availability Statement: Not applicable.
Conflicts of Interest: The authors declare no conflict of interest.
References
1.
Population Division, Department of Economic and Social Affairs, United Nations. World Population Prospects 2022: Summary
of Results. UN DESA/POP/2022/TR/NO. 3. 2022. Available online: https://www.un.org/development/desa/pd/content/
World-Population-Prospects-2022 (accessed on 10 January 2023).
2.
FAO. The Future of Food and Agriculture—Alternative Pathways to 2050; Food and Agriculture Organization of the United Nations
Rome: Rome, Italy, 2018; p. 224.
3. Umetsu, N.; Shirai, Y. Development of novel pesticides in the 21st century. J. Pestic. Sci. 2020,45, 54–74. [CrossRef]
4.
Qu, R.Y.; He, B.; Yang, J.F.; Lin, H.Y.; Yang, W.C.; Wu, Q.Y.; Li, Q.X.; Yang, G.F. Where are the new herbicides? Pest Manag. Sci.
2021,77, 2620–2625. [CrossRef] [PubMed]
5.
Heap, I. The International Herbicide-Resistant Weed Database. Online. Sunday, 27 November 2022. Available online: www.
weedscience.org (accessed on 10 January 2023).
6.
Busi, R.; Goggin, D.E.; Heap, I.M.; Horak, M.J.; Jugulam, M.; Masters, R.A.; Napier, R.M.; Riar, D.S.; Satchivi, N.M.; Torra, J.; et al.
Weed resistance to synthetic auxin herbicides. Pest Manag. Sci. 2018,74, 2265–2276. [CrossRef]
7.
Tan, X.; Calderon-Villalobos, L.I.; Sharon, M.; Zheng, C.; Robinson, C.V.; Estelle, M.; Zheng, N. Mechanism of auxin perception by
the TIR1 ubiquitin ligase. Nature 2007,446, 640–645. [CrossRef] [PubMed]
8.
Grossmann, K. Auxin herbicides: Current status of mechanism and mode of action. Pest Manag. Sci.
2010
,66, 113–120. [CrossRef]
9.
True, J.H.; Shaw, S.L. Exogenous Auxin Induces Transverse Microtubule Arrays Through Transport Inhibitor Response1/Auxin
Signaling F-Box Receptors. Plant Physiol. 2020,182, 892–907. [CrossRef] [PubMed]
10.
Calderon Villalobos, L.I.; Lee, S.; De Oliveira, C.; Ivetac, A.; Brandt, W.; Armitage, L.; Sheard, L.B.; Tan, X.; Parry, G.; Mao, H.;
et al. A combinatorial TIR1/AFB-Aux/IAA co-receptor system for differential sensing of auxin. Nat. Chem. Biol.
2012
,8, 477–485.
[CrossRef]
11.
Hoyerova, K.; Hosek, P.; Quareshy, M.; Li, J.; Klima, P.; Kubes, M.; Yemm, A.A.; Neve, P.; Tripathi, A.; Bennett, M.J.; et al. Auxin
molecular field maps define AUX1 selectivity: Many auxin herbicides are not substrates. New Phytol.
2018
,217, 1625–1639.
[CrossRef]
Molecules 2023,28, 1431 20 of 20
12.
Prigge, M.J.; Greenham, K.; Zhang, Y.; Santner, A.; Castillejo, C.; Mutka, A.M.; O’Malley, R.C.; Ecker, J.R.; Kunkel, B.N.; Estelle, M.
The Arabidopsis Auxin Receptor F-Box Proteins AFB4 and AFB5 Are Required for Response to the Synthetic Auxin Picloram. G3
(Bethesda) 2016,6, 1383–1390. [CrossRef]
13.
Schmitzer, P.R.; Balko, T.W.; Daeuble, J.F.; Epp, J.B.; Satchivi, N.M.; Siddall, T.L.; Weimer, M.R.; Yerkes, C.N. Discovery and SAR of
Halauxifen Methyl: A Novel Auxin Herbicide. In Discovery and Synthesis of Crop Protection Products; ACS Symposium Series;
American Chemical Society: Washington, DC, USA, 2015; pp. 247–260.
14.
Epp, J.B.; Alexander, A.L.; Balko, T.W.; Buysse, A.M.; Brewster, W.K.; Bryan, K.; Daeuble, J.F.; Fields, S.C.; Gast, R.E.; Green,
R.A.; et al.
The discovery of Arylex active and Rinskor active: Two novel auxin herbicides. Bioorg. Med. Chem.
2016
,24, 362–371.
[CrossRef]
15.
Zotz, A.; Bernhard, U.; Bonin, J. BELKAR
™
—A new herbicide for the control of a wide range of broadleaf weeds in winter oilseed
rape applied post-emergence in autumn. Julius-Kühn-Archiv 2018,458, 345–349. [CrossRef]
16.
Ludwig-Muller, J.; Rattunde, R.; Rossler, S.; Liedel, K.; Benade, F.; Rost, A.; Becker, J. Two Auxinic Herbicides Affect Brassica
napus Plant Hormone Levels and Induce Molecular Changes in Transcription. Biomolecules
2021
,11, 1153. [CrossRef] [PubMed]
17.
Mithila, J.; Hall, J.C.; Johnson, W.G.; Kelley, K.B.; Riechers, D.E. Evolution of Resistance to Auxinic Herbicides: Historical
Perspectives, Mechanisms of Resistance, and Implications for Broadleaf Weed Management in Agronomic Crops. Weed Sci.
2017
,
59, 445–457. [CrossRef]
18.
Yang, Z.; Li, Q.; Yin, J.; Liu, R.; Tian, H.; Duan, L.; Li, Z.; Wang, B.; Tan, W.; Liu, S. Design, synthesis and mode of action of novel
3-chloro-6-pyrazolyl picolinate derivatives as herbicide candidates. Pest Manag. Sci. 2021,77, 2252–2263. [CrossRef]
19.
Faria, J.V.; Vegi, P.F.; Miguita, A.G.C.; Dos Santos, M.S.; Boechat, N.; Bernardino, A.M.R. Recently reported biological activities of
pyrazole compounds. Bioorg. Med. Chem. 2017,25, 5891–5903. [PubMed]
20.
Havrylyuk, D.; Roman, O.; Lesyk, R. Synthetic approaches, structure activity relationship and biological applications for
pharmacologically attractive pyrazole/pyrazoline-thiazolidine-based hybrids. Eur. J. Med. Chem.
2016
,113, 145–166. [CrossRef]
21.
Huang, D.; Liu, A.; Liu, W.; Liu, X.; Ren, Y.; Zheng, X.; Pei, H.; Xiang, J.; Huang, M.; Wang, X. Synthesis and insecticidal activities
of novel 1H-pyrazole-5-carboxylic acid derivatives. Heterocycl. Commun. 2017,23, 455–460. [CrossRef]
22.
Wu, J.; Song, B.A.; Hu, D.Y.; Yue, M.; Yang, S. Design, synthesis and insecticidal activities of novel pyrazole amides containing
hydrazone substructures. Pest Manag. Sci. 2012,68, 801–810.
23.
Zhang, X.; Wei, Z.; Wang, Y.; Yang, L.; Yuan, H.; Feng, J.; Gao, Y.; Lei., P.; Ma, Z. Synthesis and antifungal activity of 3-
(difluoromethyl)-1-methyl pyrazole-4-carboxylic oxime esters. Chin. J. Pestic. Sci. 2022,24, 59–65. [CrossRef]
24.
Kim, E.; James, M.; Zhu, Y.; Gregory, T.; Christian, T.; Cynthia, A. Process for the Preparation of 4-Amino-3-chloro-5-fluoro-6-
(substituted) Picolinates. U.S. Patent 201201190857 A1, 24 January 2012.
25.
Chen, J.; Xia, Y.; Lee, S. Coupling of amides with ketones via C–N/C–H bond cleavage: A mild synthesis of 1,3-diketones. Org.
Chem. Front. 2020,7, 2931–2937. [CrossRef]
26.
Fustero, S.; Sanchez-Rosello, M.; Barrio, P.; Simon-Fuentes, A. From 2000 to mid-2010: A fruitful decade for the synthesis of
pyrazoles. Chem. Rev. 2011,111, 6984–7034. [PubMed]
27.
Zhang, Y.Z.; Tan, X.L.; Xing, W.L.; Zhang, Z. A Preparation Method for 4-Amino-3,5,6-Trichloro-2-Picolinic Acid. CN Patent
101565400 A, 5 May 2009.
28.
Lei, K.; Li, P.; Yang, X.F.; Wang, S.B.; Wang, X.K.; Hua, X.W.; Sun, B.; Ji, L.S.; Xu, X.H. Design and Synthesis of Novel 4-Hydroxyl-
3-(2-phenoxyacetyl)-pyran-2-one Derivatives for Use as Herbicides and Evaluation of Their Mode of Action. J. Agric. Food Chem.
2019,67, 10489–10497. [CrossRef] [PubMed]
29.
Kim, J.-H.; Jeong, J.-H. Structure-Activity Relationship Studies Based on 3D-QSAR CoMFA/CoMSIA for Thieno-Pyrimidine
Derivatives as Triple Negative Breast Cancer Inhibitors. Molecules 2022,27, 7974. [CrossRef] [PubMed]
Disclaimer/Publisher’s Note:
The statements, opinions and data contained in all publications are solely those of the individual
author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to
people or property resulting from any ideas, methods, instructions or products referred to in the content.