Content uploaded by Fernando Tapia
Author content
All content in this area was uploaded by Fernando Tapia on Aug 02, 2021
Content may be subject to copyright.
Caryologia. International Journal of Cytology, Cytosystematics and Cytogenetics 73(4): 17-26, 2020
Firenze University Press
www.fupress.com/caryologia
ISSN 0008-7114 (print) | ISSN 2165-5391 (online) | DOI: 10.13128/caryologia-949
Caryologia
International Journal of Cytology,
Cytosystematics and Cytogenetics
Citation: F. Tapia-Pastrana, A. Delga-
do-Salinas (2020) First cytogenetic register
of an allopolyploid lineage of the genus
Aeschynomene (Leguminosae, Papil-
ionoideae) native to Mexico. Caryolo-
gia 73(4): 17-26. doi: 10.13128/caryolo-
gia-949
Received: May 24, 2020
Accepted: November 10, 2020
Published: May 19, 2021
Copyright: © 2020 F. Tapia-Pastrana,
A. Delgado-Salinas. This is an open
access, peer-reviewed article pub-
lished by Firenze University Press
(http://www.fupress.com/caryologia)
and distributed under the terms of the
Creative Commons Attribution License,
which permits unrestricted use, distri-
bution, and reproduction in any medi-
um, provided the original author and
source are credited.
Data Availability Statement: All rel-
evant data are within the paper and its
Supporting Information les.
Competing Interests: The Author(s)
declare(s) no conict of interest.
ORCID
FTP: 0000-0003-0232-2110
First cytogenetic register of an allopolyploid
lineage of the genus Aeschynomene
(Leguminosae, Papilionoideae) native to Mexico
F T-P,*, A D-S
1 Facultad de Estudios Superiores Zaragoza, Universidad Nacional Autónoma de México,
Laboratorio de Genecología, Batalla 5 de Mayo s/n esquina Fuerte de Loreto, Col. Ejér-
cito de Oriente, Iztapalapa, C.P. 09230, Ciudad de México, Mexico
2 Instituto de Biología, Departamento de Botánica, Universidad Nacional Autónoma de
México, Apartado Postal 70-233, 04510, Cd. de México, Mexico
*Corresponding author. E-mail: pasfer@unam.mx
Abstract. A conventional cytogenetics analysis revealed for rst time an allopolyploid
lineage of the genus Aeschynomene in Mexico. e hybrid condition is conrmed aer
all the prometaphase and metaphase nuclei of the hybrids exhibited only one pair of
SAT-chromosomes, conrming the existence of nucleolar dominance and amphiplasty.
e karyotype formula for this lineage was 2n = 4x = 40 = 34 m + 6 sm with a total
diploid chromosome length (TDCL) = 28µm and an average chromosome size (AC) =
1.40 µm. Comparison of the karyotype and other chromosomal parameters with recent
cytogenetics records for other species of the subgenus Aeschynomene included in the
Nod-independent clade allows propose to Aeschynomene evenia and A. scabra as possi-
ble progenitors. Furthermore, other comparison of seedlings focused at the number of
leaets of the rst four eophylls of the proposed parents and of the hybrid individuals
allowed to observe coincidences that support the proposal made from the cytogenetic
analysis. Evidence of “gigas” eects on owers and fruits of hybrids is also shown.
Keywords: cryptic taxa, cytotype, karyotype, nucleolar dominance, SAT-chromo-
somes, secondary constrictions, seedlings.
I. INTRODUCTION
Aeschynomene Linnaeus (Leguminosae, Tribe Dalbergieae s. l.) is a
diverse genus of subfamily Papilionoideae (Papilionoid legumes) distributed
in the tropics and subtropics of the world (Lavin et al. 2001, Klitgaard and
Lavin 2005). It comprises herbaceous and woody species, annual, repetitive
and perennial with dierent ecological requirements. Several species con-
tribute to supplement nitrogen to the soil through the production of nodular
roots and stems in symbiosis with nitrogen xing bacteria, so they are eco-
nomically important as green manure (Alazar and Becker 1987; Fernandes
1996; Souza et al. 2012) and recently, Aeschynomene evenia C. Wright has
been proposed as a model species in genetics to develop new agronomic
18 Fernando Tapia-Pastrana, Alfonso Delgado-Salinas
strategies in the engineering of nitrogen xing nodules
that enhance rice production (Arrighi et al. 2012, 2013).
is taxon belongs to the group of 11 semi-aquatic spe-
cies of Aeschynomene that have the property of being
nodulated by photosynthetic Bradyrhizobium that lack
the nodABC genes necessary for the synthesis of Nod
factors and are grouped into the so-called Nod-inde-
pendent clade (Chaintreuil et al. 2013; Brottier et al.
2018) and that correspond to the morphological series
Indicae and Sensitivae (Rudd 1955).
e genus Aeschynomene traditionally included in
the Aeschynomeneae tribe (Polhill et al.1981) and cur-
rently circumscribed in the Dalbergioid clade (Lavin et
al. 2001; Wojciechowski et al. 2004) has evolved in dif-
ferent ecological niches and includes herbaceous forms,
annual and perennial shrubs and trees up to 8 meters,
with compound pinnate leaves and papilionoid owers
that are generally self-pollinated, although there is cross-
pollination by bees (Rudd 1955; Fernandes 1996; Arrighi
et al. 2014, Carleial et al. 2015). Other studies indicate
that the genus Aeschynomene is not monophyletic and
taxa with basixed stipules and a campanulate calyx
(subgenus Ochopodium Vogel) are more related to the
genera Machaerium Persoon and Dalbergia Linnaeus f.
than to taxa with medxed stipules and a bilabiate calyx
(subgenus Aeschynomene Léonard) (Ribeiro et al. 2007;
Cardoso et al. 2012).
Currently Aeschynomene genus contains 170 (http://
www.theplantlist.org) to 180 species (Klitgaard and
Lavin 2005) 231taxa and cytotypes at four ploidy levels:
diploid (2x), tetraploid (4x), hexaploid (6x) and octoploid
(8x) (Index to Plant Chromosome Numbers; Kawakami
1930; Bielig 1997; Arrighi et al. 2012, 2014; Chaintreuil
et al. 2016, 2018; Brottier et al. 2018). America, where
most of the taxa are 2n = 20 diploids, has been proposed
as the center of origin of the genus, with a secondary
distribution in Africa and Asia where polyploid species
and some cases of aneuploidy predominate (Chaintreuil
et al. 2018; Tapia-Pastrana et al. 2020).
Although it is clear that in the Dalbergioid clade,
diploid 2n = 20 genera predominate, with some poly-
ploid and aneuploid species, in Aeschynomene there is
currently a renewed interest in knowing to what extent
polyploidy has contributed to the diversication and
radiation of the group. In this respect Arrighi et al.
(2014) revealed multiple hybridization/polyploidiza-
tion events, highlighting the prominent role of allopoly-
ploidy in the diversication of Nod-independent clade.
In addition Chaintreuil et al. (2016) studied African
Aeschynomene species and their data support the idea
that the whole African group is fundamentally tetraploid
and revealed the allopolyploid origin of A. afraspera
J. Léonard (2n = 8x = 76) and A. schimperi Hochst. ex
A.Rich. (2n = 8x = 56), where variations in the number
of chromosomes also indicated possible dysploidy/ane-
uploidy events. In Mexico, Aeschynomene is represented
by 31 species and infraspecic taxa including several
endemisms. An investigation about the patterns of chro-
mosomal evolution in Mexican species, including six
taxa of the Nod-independent clade, showed the predomi-
nant of a basic 2n = 20 diploid structure and evolution-
ary patterns related to the corresponding morphological
series (Tapia-Pastrana et al. 2020).
In the present research, a conventional cytogenetic
study was carried out to obtain the karyotype and ana-
lyze the level of ploidy in a Mexican population initially
described as Aeschynomene scabra G. Don, where the
size of the owers, fruits and seeds generated suspicions
about a possible hybrid origin. In addition as the sam-
pled individuals exhibited oral morphotypes similar to
those of A. evenia C. Wright and A. scabra, whose col-
lection records in Mexico would support their partici-
pation in the hybridization process, the growth pattern
of the rst four eophylls was also compared in putative
hybrids and their parental assumptions.
2. MATERIAL AND METHODS
2.1 Collection sites
Seeds of the putative hybrids were collected in the
Municipio de la Huerta, Estado de Jalisco, Mexico,
19°29´N; 105°01´W (Carleial s/n, MEXU). e climate
is semi-dry and warm. Mean temperature in the area is
25.2 °C, and there is a well-dened rainy season (average
annual precipitation: 1107mm) occurring from June to
October (García-Oliva et al. 20 02).
e seeds of Aeschynomene evenia and A. scabra
were collected in the municipalities of Coyuca de Cat-
alán (18° 19´ N; 100° 42´ W, JC Soto 15333 (MEXU))
and Arcelia (18°18´54´´N; 100°17´02´´W, JC Soto 15393
(MEXU)) respectively, in the State of Guerrero, Mexi-
co. Both municipalities are part of the Tierra Caliente
region. e predominant climate is warm subhumid
with rains from June to September (average annual
precipitation: 1100 to 1200 mm). e studied taxa are
assigned to the infrageneric classication of Neotropi-
cal Aeschynomene sensu Rudd (1955) series Indicae of
subgenus Aeschynomene and are part of the Nod-inde-
pendent monophyletic clade (Chaintreuil et al. 2013),
whose taxa are nodulated on roots and stems by photo-
synthetic Bradyrhizobium strains lacking the nod ABC
genes necessary for the synthesis of Nod factors (Giraud
et al. 2007).
19
First cytogenetic register of an allopolyploid lineage of the genus Aeschynomene native to Mexico
2.2 Chromosome and karyotype procedures in putative
hybrids
Seeds were collected in summer 2014 and from at
least six plants. Batches of 40 seeds from each plant were
used. e seeds were scaried and germinated in Petri
dishes lined with a moist lter paper at room tempera-
ture and under natural light. Chromosomes at metaphase
and prophase were obtained following the splash method
(Tapia-Pastrana and Mercado-Ruaro 2001). All meris-
tems were collected from 2-4 mm long roots pretreated
with 2 mM 8-hydroxyquinolin for 5 h at room tempera-
ture and xed in the xative (ethanol: acetic acid=3:1).
ey were then treated with a mixture of 20% pectinase
(Sigma) and 2% cellulase (Sigma) in 75 mM KCl for 60
min at 37 °C. Aer centrifugation at 1500 rpm for 10
min, the cell pellet was transferred to 75 mM KCl solu-
tion for 13 min at 37 °C. Aer two successive rinses with
the KCl solution, they were again xed in the xative and
subsequently rinsed twice more. One or two drops of the
suspension of pellet were placed on clean slides, air-dried
and stained in 10% Giemsa for 13 min. Preparations were
made permanent using a synthetic resin.
At least ten metaphase plates of intact cells with
well-spread chromosomes, no chromosome overlap-
ping, and same contraction and ten prophase plates were
photographed from each collection, using a microscope
(Axioscope, Carl Zeiss) and analyzed for chromosome
number determinations. Five photographs of meta-
phases with chromosomes having similar comparable
degrees of contraction and centromeres clearly located
were utilized to obtain the Total diploid chromosome
length (TDCL), Total chromosome length (TCL), Aver-
age chromosome length (AC), the dierence in length
between the longest chromosome and the shortest chro-
mosome (Range) and the longest/shortest chromosome
ratio (L/S). e shapes of chromosomes were classied
according to Levan et al. (1964) and the TF was obtained
following Huziwara (1962). Furthermore, prometaphase
cells were analyzed to verify both the number of nucleo-
li, and the behavior of the SAT chromosomes. e infor-
mation thus obtained was compared with that recently
recorded for Aeschynomene evenia and A. scabra in
another cytogenetic study where the same method was
used for karyotype analysis in Aeschynomene species
and varieties (Tapia-Pastrana et al. 2020).
2.3 Seedlings and Eophylls
In order to compare seedling morphology in indi-
viduals of the supposedly hybrid population with those
of Aeschynomene evenia and A. scabra, the development
of 20 individuals grown in pots under greenhouse con-
ditions was evaluated. Interest was particularly focused
on the number of leaets and the presence of hairs on
their edges until the complete development of the fourth
leaf. Eophylls at the first, second, third and fourth
eophyllar nodes were referred to as E1, E2, E3 and E4,
respectively following Schütz et al. (2019). Photographs
of seedlings were taken with a Canon SX700 HS camera.
3. RESULTS
3.1 Karyotype analysis
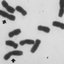
A total of 410 cells were analyzed in metaphase
and 16 in prometaphase and all exhibited a 2n = 4x =
40 (Fig. 1 A-C). TDCL was 28 µm and AC 1.40 µm. e
chromosomal range was 0.56 µm, the ratio 1.48 and
a TF = 42.46. e karyotype formula was 2n = 4x =
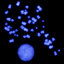
34m + 6sm (Table 1). Consistently, in all prometaphase
and metaphase nuclei, only one pair of submetacentric
chromosomes was observed having lax secondary con-
strictions and macrosatellites in short arms (SAT-chro-
mosomes) (Fig. 1 A-C). e karyotype exhibits small
chromosomes (1.72-1.16 µm) clearly discernible, with
predominance of metacentric chromosomes (m) and
lacking subtelocentric chromosomes (st). is arrange-
ment is consistent with a TF that describes a slightly
asymmetric karyotype (Fig. 1D and Table 1). Occasion-
ally the SAT-chromosomes were observed immersed in a
single nucleolus.
3.2 Seedlings and Eophylls
e seedlings of the three taxa are illustrated in Fig.
2 A-C. Eophylls are stipulated, alternate, petiolate, pin-
nate, with alternate leaets, have elliptic to oblong leaf-
lets, a rounded apex, an entire margins, and one cen-
tral primary vein in the three taxa under study. e
leaflets did not present trichomes; both adaxial and
abaxial surfaces are glabrous. e number of leaets
in the rst four eophylls in seedlings of individuals of
Aeschynomene evenia, A. scabra and putative hybrids are
shown in Tables 2-4 respectively.
4. DISCUSSION
It is clear that the entire Dalbergioid clade
(Adesmia, Dalbergia and Pterocarpus subclades) is dom-
inated by 2n = 2x = 20 species, with scattered polyploids
and aneuploids (Lavin et al. 2001). In addition an ances-
20 Fernando Tapia-Pastrana, Alfonso Delgado-Salinas
tral state reconstruction performed in a phylogeny based
on ITS + matK of the Aeschynomene genus and related
genera indicated that diploidy is the ancestral condi-
tion in the entire group reviewed (Brottier et al. 2018).
However, the role of allopolyploid speciation events in
the origin of new taxa is now recognized (Arrighi et al.
2014).
As far as we know, the rst assumption about of
hybridization in Aeschynomene is attributed to Rudd
(1955) who pointed out that the species with the widest
distribution within the Indicae series (Nod-independent
clade) tend to be more variable and intergrade with their
neighbors. Later, Verdcourt (1971) suggested that speci-
mens of Aeschynomene rudis Bentham (also into Nod-
independent clade) with large owers could be of poly-
ploid origin, without pointing out the possible duplica-
tion mechanism involved, auto or allopolyploidy. To
date, several studies have shown that the clade of A. eve-
nia is mainly diploid (2n = 2x = 20), however some spe-
cies such as A. indica Linnaeus (2n = 4x = 40, 2n = 6x
= 60) seem to be of recent allopolyploid origin (Arrighi
et al. 2014; Chaintreuil et al. 2018; Tapia-Pastrana et al.
2020). Furthermore, it has been found that all species of
the group A. afraspera are polyploid (2n = 4x = 28, 38,
40; 2n = 8x = 56, 76) and have a common AB genomic
structure (Chaintreuil et al. 2016). In facts phylogenetic
relationships between diploids and polyploids elucidated
from ITS sequences show that in the Nod-independent
clade, species such as A. evenia, A. scabra and A. rudis
participate in the hybridization/polyploidization events
and formation of polyploid complexes that have contrib-
uted to the radiation of this group (Arrighi et al. 2014).
Figure 1. Mitotic metaphase cells of hybrid Aeschynomene 2n = 4x
= 40. A-C, Metaphase chromosome plates in optimal spread; D,
Karyotype 34m + 6sm. e chromosomes are aligned in decreas-
ing order. Arrows point to secondary constrictions and satellites on
short arms of submetacentric chromosomes.
Table 1. Average chromosome measurements obtained from ve
nuclei in metaphase of the hybrid population (2n = 4x = 40 = 34m
+ 6sm) under study.
CP TCL
(µm)
LLA
( µm)
LSA
(µm) r CT
01 1.72 0.96 0.77 1.24 m
02 1.63 0.89 0.73 1.21 m
03 1.59 0.89 0.69 1.28 m
04 1.55 0.81 0.72 1.12 m
05 1.53 0.81 0.70 1.15 m
06 1.50 0.82 0.66 1.24 m
07 1.48 0.83 0.63 1.31 m
08 1.46 0.79 0.66 1.19 m
09 1.44 0.79 0.63 1.25 m
10 1.42 0.79 0.61 1.29 m
11 1.38 0.74 0.64 1.17 m
12 1.36 0.89 0.46 1.93 sm*
13 1.34 0.78 0.55 1.41 m
14 1.31 0.70 0.59 1.18 m
15 1.28 0.72 0.55 1.30 m
16 1.24 0.83 0.40 2.07 sm
17 1.23 0.69 0.52 1.32 m
18 1.19 0.67 0.54 1.24 m
19 1.19 0.66 0.49 1.34 m
20 1.16 0.78 0.36 2.16 sm
TDCL 28.00
AC 1.40
Abbreviations: CP- chromosome pair; TCL- total chromosome
length; LLA- length long arm; LSA- length short arm; r- arm ratio;
CT-chromosome type; TDCL- Total diploid chromosome length;
AC- Average chromosome length; m- metacentric; sm- submeta-
centric; *- satellite. Abbreviations: CP- chromosome pair; TCL-
total chromosome length; LLA- length long arm; LSA- length short
arm; r- arm ratio; CT-chromosome type; TDCL- Total diploid chro-
mosome length; AC- Average chromosome length; m- metacentric;
sm- submetacentric; *- satellite.
21
First cytogenetic register of an allopolyploid lineage of the genus Aeschynomene native to Mexico
In the present investigation, the chromosomal num-
ber obtained in all the nuclei analyzed from the individ-
uals under study was 2n = 4x = 40, which undoubtedly
shows that they are polyploid cells and that the individ-
uals from which they come integrate a polyploid lineage
not previously detected in Mexico (Rudd 1955; Tapia-
Pastrana et al. 2020). e origin of the polyploidy (auto
or allopolyploidy) were established easily from the num-
ber of SAT chromosomes unambiguously identied both
in nuclei in prometaphase and metaphase and by their
position in relation to the nucleolus.
Indeed, polyploidy, the process of genome dou-
bling that gives rise to organism with multiple sets of
chromosomes, is recognized as an important process in
plant evolution, a major mechanism of adaptation and is
oen invoked as a driver of diversication (Ramsey and
Figure 2. Seedling morphology of Aeschynomene under study until the complete development of the fourth eophyll A, Aeschynomene eve-
nia; B, A. scabra; C, hybrid of Aeschynomene.
Table 2. Number of leaets up to the fourth eophyll in Aeschynomene evenia.
1 2 3 4 5 6 7 8 9 10 11 12 13 14 15 16 17 18 19 20
E1 10 10 10 10 10 10 10 10 10 10 10 9 10 9 8 10 9 10 10 8
E2 12 12 12 12 11 12 14 12 10 14 12 12 14 12 10 13 12 11 10 11
E3 15 16 15 14 14 16 17 15 14 16 16 15 17 12 12 16 16 12 14 12
E4 16 18 16 16 16 16 18 18 14 16 18 16 18 16 15 16 16 14 16 15
Table 3. Number of leaets up to fourth eophyll in Aeschynomene scabra.
1 2 3 4 5 6 7 8 9 10 11 12 13 14 15 16 17 18 19 20
E1 10 8 10 10 10 10 10 8 10 9 10 10 10 10 10 8 8 8 8 10
E2 12 15 14 14 14 14 14 12 16 13 14 13 14 14 14 12 12 13 15 14
E3 20 20 22 18 18 18 19 19 20 17 17 18 18 16 18 19 19 19 20 16
E4 24 25 27 23 22 22 22 22 24 22 22 22 22 21 22 22 22 20 25 21
Table 4. Number of leaets up to fourth eophyll in hybrids.
1 2 3 4 5 6 7 8 9 10 11 12 13 14 15 16 17 18 19 20
E1 10 10 8 10 10 9 10 8 8 10 8 10 8 10 10 8 10 10 8 10
E2 14 14 12 14 13 14 14 14 14 14 14 14 14 16 14 14 14 14 14 10
E3 18 20 18 20 20 19 20 20 20 19 18 20 18 20 20 20 20 15 18 14
E4 21 25 22 23 23 26 23 24 23 24 23 26 22 27 22 22 23 22 23 14
22 Fernando Tapia-Pastrana, Alfonso Delgado-Salinas
Schemske 1998; Soltis et al. 2009) and it is likely to be
one of the most predominant mechanisms of sympat-
ric speciation in plants (Otto and Whitton 2000). It can
act alone, resulting in autopolyploidy, or in concert with
hybridization, producing allopolyploids, and both modes
lead to plant speciation. It should be mentioned that in
the process of polyploidization by total gene duplica-
tion (autopolyploidy) the number of satellites present
in a diploid species is also doubled, since this does not
involve loss or suppression of the nucleolar function, the
NOR regions associated with secondary constrictions
in SAT-chromosomes are they show lax and therefore
satellites are clearly appreciated. It is known that NORs
contain tandemly arranged highly reiterated riboso-
mal rRNA genes coding for 18S-5.8S-26S rRNA whose
expression is under epigenetic control (Pikaard 2000).
For example, Medicago sativa Linnaeus, a recognized
autotetraploid exhibits four macrosatélites in metaphase
cells (Falistocco 1987). In contrast, plants of allopoly-
ploid origin as cotton (Gossypium hirsutum Linnaeus
2n = 4x = 52 AADD, Endrizzi et al. 1985), wheat (Triti-
cum aestivum Linnaeus 2n = 6x = 42, AABBDD, Laca-
dena and Cermeño 1985; Friebre et al. 1995) and canola
(Brassica napus Linnaeus 2n = 4x = 38 AACC; Xiong
and Pires 2011) undergo inactivation of the regions of
the nucleolar organizer (NOR) of one of the parental
genomes, silenced by the eect of nucleolar dominance
(Navashin 1934) and consequently a smaller num-
ber of satellites is recorded (Doyle et al. 2008; Ge et al.
2013). It is, rDNA loci may be additive in number, but
then exhibit dierences in gene expression. Interspecic
hybrids oen have rRNA genes of one parent function-
ally dominant over the rRNA of the other parent, and
there are many examples of such regulation of rRNA
gene activity in allopolyploids (Pikaard 2000; Pires et
al. 2004). Comparative analyses of nucleolar organizer
regions (NORs) of somatic metaphase chromosomes
made by phase contrast, C-banding and silver staining
have demonstrated that the activity of the NORs of cer-
tain chromosomes can be suppressed or partially inhib-
ited by the presence of other SAT-chromosomes.
e NOR competition is cytologically expressed as
amphiplasty: a term proposed to denote morphological
changes which occur in chromosomes following inter-
specic hybridization (Rieger et al. 1976). e second-
ary constriction of the SAT-chromosome of one of the
parental species is missing in the hybrid and the satellite
is retracted onto the chromosome arm as a consequence
(Lacadena and Cermeño 1985). us, in the Hordeum
murinum Linnaeus complex (Poaceae, Triticeae), tetra-
ploid and hexaploid cytotypes arising from hybridiza-
tion exhibit only a pair of chromosomes with second-
ary and satellite constrictions (Cuadrado et al. 2013). In
fact, the inactivation or epigenetic silencing of ribosomal
genes is one of the most common phenomena in hybrid
and polyploid members of Triticeae Linnaeus (Cermeño
and Lacadena 1985; Carmona et al. 2016) and one of the
rst examples of dierential gene expression discovered
in plant hybrids nearly a century ago (Navashin 1934;
Matyásӗk et al. 2007). In the present work, the repres-
sive eects on NORs from allopolyploid population are
cytologically expressed (amphiplasty) as the suppression
of a secondary constriction clearly observed in all their
complements (Fig. 1 A-D).
e karyotype exhibited in hybrid individuals (34m
+ 6sm) (Fig. 1D and Table 1) coincides in several respects
with that expected at a cross between A. evenia (2n =
2x = 7m + 3sm) and A. scabra (2n = 2x = 10 m) (Fig. 2
in Tapia-Pastrana et al. 2020). For example, the num-
ber of sm chromosomes in A. evenia agrees with the
6sm in hybrid individuals. In addition to submetacen-
tric chromosomes, these individuals exhibit metacentric
chromosomes whose predominance is consistent with
the karyotype formulas described in their putative rela-
tives, whose complements lack subtelocentric chromo-
somes (Tapia-Pastrana et al. 2020). ere is a coincidence
between THC and AC and even the morphology of the
SAT-chromosomes (submetacentrics with macrosatellites
in short arms) and their position in the karyotype is very
similar to that recently described in A. scabra (Tapia-
Pastrana et al. 2020). erefore we propose to A. evenia
and A. scabra as progenitors of the allopolyploid popula-
tion (2n = 4x = 40 = 34m + 6sm) registered in this work.
e reasoning is simple: if a diploid species is involved
in the origin of a tetraploid cytotype, its chromosomes
must be present in it. e same is true if tetraploid forms
are involved in the origin of hexaploid forms (Cuadrado
et al. 2013). In Mexico, recent collection data shows that
populations of both species occupy overlapping ranges in
some central areas of the country where A. evenia is con-
sidered an introduced species (Arrighi et al. 2013; Chain-
treuil et al. 2018; Tapia-Pastrana et al. 2020).
is new proposal is not surprising, since previously
the Indicae series species grouped within Nod-inde-
pendent clade, including A. evenia and A. scabra, have
been identied as progenitors in allopolyploids and in
the formation of polyploid complexes, although attempts
at hybridization have failed to form fertile individu-
als (Arrighi et al. 2014). Regarding the identity of the
allopolyploid taxon recorded here, it can be argued that
a detailed review of its complete morphological charac-
ters (data not shown) suggests that it shares character-
istics described for Aeschynomene rudis particularly in
the shape and size of owers, fruits (hispidulous, verru-
23
First cytogenetic register of an allopolyploid lineage of the genus Aeschynomene native to Mexico
coses, or muricate at the center) and seeds (Rudd 1955).
However, it also recalls the robust version of A. scabra
described by Rudd (1955). e existence of cryptic taxa
in Aeschynomene as well as the need for broader sam-
pling to detect new cytotypes has already been pointed
out (Brottier et al. 2018, Chaintreuil et al. 2018) and the
results of this study conrm this.
Regarding the results obtained from the seedling
comparison, these seem to support a close relationship
between the individuals of the three populations studied
(Fig. 2, Tables 2-4). In principle, the observed intervals
in the number of leaets per eophyll (E1-E4) show some
uniformity, particularly E1, whose interval (8-10 leaf-
lets) was repeated in the three populations. In interme-
diate eophylls (E2-E4) a close concordance is observed
between A. scabra and the hybrid population, while
in A. evenia the number of leaets was lower in corre-
spondence with the taxonomic description of this spe-
cies (Rudd 1955). Furthermore, the morphology of the
eophylls was similar and in all populations the leaets
Figure 3. Floral morphotypes (above), dissected owers and fruits (below) of the taxa under study. A, D and E, Aeschynomene evenia; B, F
and G, A. scabra; C, H and I, hybrid of Aeschynomene. All three taxa exhibit typical pea or papilionoid owers. ese zygomorphic owers
comprise a standard (vexillum or banner) petal (adaxially placed), two lateral petals (wings) and two (usually partially fused and abaxially
placed) keel petals, which conceal the androecium and gynoecium. e fruits have similar characteristics and are mainly dierentiated by
their size. Above scale bar = 0.5 cm, below = 1.0 cm.
24 Fernando Tapia-Pastrana, Alfonso Delgado-Salinas
exhibited entire margins, without trichomes and with a
central primary vein.
Polyploids are known to oen have novel pheno-
types that are not present in their diploid progenitors or
that exceed the range of parent species (“gigas” eects)
(Ramsey and Schemske 2002; Ramsey and Ramsey 2014).
In this sense, Fig. 3 shows oral morphotypes, dissected
owers and fruits of the populations studied here, where
similarities are observed, but the dierences in size of
such characters are highlighted. e results obtained in
this study conrm that in the Nod-independent lineage
within the genus Aeschynomene, hybridization and poly-
ploidization play a relevant role in the formation of spe-
cies and those taxa such as the polymorphics A. evenia
and A. scabra actively participate in it.
ACKNOWLEDGEMENTS
is study is part of the doctoral thesis of the rst
author, F T-P, carried out at the Posgrado en Ciencias
Biológicas of the Universidad Nacional Autónoma de
México (UNAM). e authors thank to Dr. Samuel Car-
leial for seeds originally identied as A. scabra and to
the Division of Postgraduate Studies and Research of the
Faculty of Higher Studies, Zaragoza, UNAM for the sup-
port provided during the development of this research.
REFERENCES
Alazar D, Becker M. 1987. Aeschynomene as green
manure for rice. Plant and Soil 101: 141-143.
Arrighi JF, Cartieaux F, Brown SC, Rodier-Goud M,
Boursot M, Fardoux J, Patrel D, Gully D, Fabre S,
Chaintreuil C, Giraud E. 2012. Aeschynomene evenia,
a model plant for studying the molecular genetics of
the Nod-independent rhizobium-legume symbiosis.
Molecular Plant-Microbe Interactions 25: 851-861.
Arrighi J-F, Cartieaux F, Chaintreuil C, Brown S, Bour-
sot M, Giraud E. 2013. Genotype delimitation in the
Nod-Independent model legume Aeschynomene eve-
nia. PLoS ONE 8(5): e63836. doi:10.1371/journal.
pone.0063836
Arrighi JF, Chaintreuil C, Cartieaux F, Cardi C, Rodier-
Goud M, Brown SC, Boursot M, D´Hont A, Dreyfus
B, Giraud E. 2014. Radiation of the Nod-independent
Aeschynomene relies on multiple allopolyploid specia-
tion events. New Phytologist 201: 1457-1468.
Bielig LM. 1997. Chromosome numbers in the forage
legume genus, Aeschynomene L. SABRAO Journal
29:33-39.
Brottier L, Chaintreuil C, Simion P, Scornavacca C, Rival-
lan R, Mournet P, Moulin L, Lewis GP, Fardoux J,
Brown SC, Gomez-Pacheco M, Bourges M, Hervouet
C, Gueye M, Duponnois R, Ramanankierana H, Ran-
driambanona H, Vandrot H, Zabaleta M, DasGupta
M, D’Hont A, Giraud E, Arrighi JF. 2018. A phyloge-
netic framework of the legume genus Aeschynomene
for comparative genetic analysis of the Nod-depend-
ent and Nod-independent symbioses. BMC Plant
Biology 18: 333.
Cardoso D, de Queiroz LP, Pennington RT, de Lima HC,
Fonty E, Wojciechowski MF, Lavin M. 2012. Revis-
iting the phylogeny of papilionoid legumes: New
insights from comprehensively sampled early-branch-
ing lineages. American Journal of Botany 99: 1991-
2013.
Carleial S, Delgado-Salinas A, Domínguez CA, Terrazas
T. 2015. Reexed owers in Aeschynomene amor-
phoides (Fabaceae: Faboideae): a mechanism promot-
ing pollination specialization? Botanical Journal of
the Linnean Society 177: 657-666.
Carmona A, de Bustos A, Jouve N, Cuadrado A. 2016.
Allopolyploidy and the complex phylogenetic rela-
tionships within the Hordeum brachyantherum taxon.
Molecular Phylogenetics and Evolution 97: 107-119.
Cermeño MC, Lacadena JR. 1985. Nucleolar organizer
competition in Aegilops–rye hybrids. Canadian Jour-
nal of Genetics and Cytology 4: 479-483.
Chaintreuil C, Arrighi JF, Giraud E, Miché L, Moulin L,
Dreyfus B, Munive-Hernández J-A, Villegas-Hernán-
dez MC, Béna G. 2013. Evolution of symbiosis in the
legume genus Aeschynomene. New Phytologist 200:
1247-1259.
Chaintreuil C, Gully D, Hervouet C, Tittabutr P, Ran-
driambanona H, Brown SC, Lewis GP, Bourge M,
Cartieaux F, Boursot M, Ramanankierana H, D´Hont
A, Teaumroong N, Giraud E, Arrighi JF. 2016. e
evolutionary dynamics of ancient and recent poly-
ploidy in the African semiaquatic species of the leg-
ume genus Aeschynomene. New Phytologist 211:
1077-1091.
Chaintreuil C, Perrier X, Guillaume M, Fardoux J, Lew-
is GP, Brottier L, Rivallan R, Gomez- Pacheco M,
Bourges M, Lamy L, ibaud B, Ramanankierana H,
Randriambanona H, Vandrot H, Mournet P, Giraud
E, Arrighi JF. 2018. Naturally occurring variations in
the nod-independent model legume Aeschynomene
evenia and relatives: a resource for nodulation genet-
ics. BMC Plant Biology 18: 54.
Cuadrado A, Carmona A, Jouve N. 2013. Chromosomal
characterization of the three subgenomes in the poly-
ploids of Hordeum murinum L.: New insight into the
25
First cytogenetic register of an allopolyploid lineage of the genus Aeschynomene native to Mexico
evolution of this complex. PLoS ONE 8(12),e81385.
doi:10.1371/journal.pone.0081385
Doyle JJ, Flagel LE, Paterson AH, Rapp RA, Soltis DE,
Soltis PS, Wendel JF. 2008. Evolutionary genetics
of genome merger and doubling in plants. Annual
Review of Genetics 42: 443-461.
Endrizzi JE, Turcotte EL, Kohel RJ. 1985. Genetics, cytol-
ogy, and evolution of Gossypium. Advances in Genet-
ics 23: 271-375.
Falistocco E. 1987. Cytogenetic investigations and karyo-
logical relationships of two Medicago: M. sativa L.
(Alfalfa) and M. arborea L. Caryologia 4: 339-346.
Fernandes A. 1996. O táxon Aeschynomene no Brasil. -
Fortaleza: Edições UFC. Brasil.
Friebre B, Jiang J, Tulee N, Gill BS. 1995. Standard karyo-
type of Triticum umbellulatum and the characteriza-
tion of derived chromosome addition and transloca-
tion lines in common wheat. eoretical and Applied
Genetics 90: 150-156.
García-Oliva F, Camou A, Maass JM. 2002. El clima de
la región central de la costa del Pacíco Mexicano.
In: Noguera, F. A., Vega-Rivera, J. H., García-Aldrete,
A. N., Quesada-Avendaño, M. eds. Historia natu-
ral de Chamela. Mexico City: Universidad Nacional
Autónoma de México, Instituto de Biología, 3-10.
Ge X-H, Ding L, Li Z-Y. 2013. Nucleolar dominance and
dierent genome behaviors in hybrids and allopoly-
ploids. Plant Cell Reports 32: 1661-1673.
Giraud E, Moulin L, Vallenet D, Barbe V, Cytryn E, Ava-
rre JC, Jaubert M, Simon D, Cartieaux F, Prin Y,
Bena G, Hannibal L, Fardoux J, Kojadinovic M, Vuil-
let L, Lajus A, Cruveiller S, Rouy Z, Mangenot S,
Segurens B, Dossat C, Franck WL, Chang W-S, Saun-
ders E, Bruce D, Richardson P, Normand P, Drey-
fus B, Pignol D, Stacey G, Emerich D, Vermeglio A,
Medigue C, Sadowsky M. 2007. Legumes symbioses:
absence of Nod genes in photosynthetic Bradyrhizo-
bia. Science 316(5829): 1307-1312.
Huziwara Y. 1962. Karyotype analysis in some genera of
Compositae. VIII. Further studies on the chromo-
somes of Aster. American Journal of Botany 49: 116-
119.
Kawakami J. 1930. Chromosome numbers in Leguminos-
ae. e Botanical Magazine, Tokyo 44: 319-328.
Klitgaard BB, Lavin M. 2005. Tribe Dalbergieae sensu
lato. In: Lewis GP, Schrire BD, Mackinder BA, Lock
JM (Eds) Legumes of the world. Royal Botanic Gar-
dens, Kew Publishing, London, 306-335.
Lacadena JR, Cermeño MC. 1985. Nucleolus organizer
competition in Triticum aestivum - Aegilops umbel-
lulata chromosome addition lines. eoretical and
Applied Genetics 71: 278-283.
Lavin M, Pennington RT, Klitgaard BB, Sprent JI, de
Lima HC, Gasson PE. 2001. e dalbergioid legumes
(Fabaceae): delimitation of a pantropical monophyl-
etic clade. American Journal of Botany 88: 503-533.
Levan A, Fredga K, Sandberg AA. 1964. Nomenclature
for centromeric position on chromosomes. Hereditas
52: 201-219.
Matyásӗk R, Tate JA, Lim YK, Šrubařová H, Koh J, Leitch
AR, Soltis DE, Soltis PS, Kovařík A. 2007. Concerted
evolution of rDNA in recently formed Tragopogon
allotetraploids is typically associated with an inverse
correlation between gene copy number and expres-
sion. Genetics 176: 2509-2519.
Navashin M. 1934. Chromosome alterations caused by
hybridization and their bearing upon certain general
genetic problems. Cytologia 5: 169-203.
Otto SP, Whitton J. 2000. Polyploid incidence and evolu-
tion. Annual Review of Genetics 34: 401-437.
Pikaard CS. 2000. e epigenetics of nucleolar domi-
nance. Trends in Genetics 16: 495-500.
Pires JC, Lim KY, Kovarík A, Matyásek R, Boyd A, Leitch
AR, Leitch IJ, Bennett MD, Soltis PS, Soltis DE. 2004.
Molecular cytogenetic analysis of recently evolved
Tragopogon (Asteraceae) allopolyploids reveal a kar-
yotype that is additive of the diploid progenitors.
American Journal of Botany 91: 1022-1035.
Polhill RM, Raven PH, Stirton CH. 1981. Evolution
and systematics of the Leguminosae. In Polhill RM,
Raven PH (eds.) Advances in Legume Systematics,
part 1, 1-26. Royal Botanic Gardens, Kew, UK.
Ramsey J, Schemske DW. 1998. Pathways, mechanisms, and
rates of polyploid formation in owering plants. Annu-
al Review of Ecology and Systematics 29: 467-501.
Ramsey J, Schemske DW. 2002. Neopolyploidy in ower-
ing plants. Annual Reviews in Ecology and Systemat-
ics 33: 589-639.
Ramsey J, Ramsey TS. 2014. Ecological studies of poly-
ploidy in the 100 years following its discovery.
Philosophical Transactions of Royal Society B 369:
20130352.
Ribeiro RA, Lavin M, Lemos-Filho JP, Mendonça Filho
CV, Rodrigues dos Santos F, Lovato MB. 2007. e
genus Machaerium (Leguminosae) is more closely
related to Aeschynomene sect. Ochopodium than to
Dalbergia: inferences from combined sequence data.
Systematic Botany 32: 762-771.
Rieger R, Michaelis A, Green MM. 1976. Glossary of
genetics and cytogenetics: classical and molecular.
Springer-Verlag, Berlin. 647 pp.
Rudd VE. 1955. e American species of Aeschynomene.
Contributions from the United States National Her-
barium 32: 1-172.
26 Fernando Tapia-Pastrana, Alfonso Delgado-Salinas
Schütz RR, da Silva HL, Silva FA. 2019. Seedling mor-
phology of some Brazilian taxa of Aeschynomene
(Leguminosae) and its systematic relevance. Flora
255: 69-79.
Soltis DE, Albert VA, LeebensMack J, Bell CD, Paterson
AH, Zheng C, Sanko D, de Pamphilis CW, Kerr Wall
P, Soltis PS. 2009. Polyploidy and angiosperm diversi-
cation. American Journal of Botany 96: 336-348.
Souza MC, Vianna LF, Kawakita K, Miotto STS. 2012. O
gênero Aeschynomene L. (Leguminosae, Faboideae,
Dalbergieae) na planicie de inundação do alto rio
Paraná, Brasil. Revista Brasileira de Biociências 10:
198-210.
Tapia-Pastrana F, Delgado-Salinas A, Caballero J. 2020.
Patterns of chromosomal variation in Mexican spe-
cies of Aeschynomene (Fabaceae, Papilionoideae) and
their evolutionary and taxonomic implications. Com-
parative Cytogenetics 14: 157-182.
Tapia-Pastrana F, Mercado-Ruaro P. 2001. A combina-
tion of the “squash” and “splash” techniques to obtain
the karyotype and asses meiotic behavior of Prosopis
laevigata L. (Fabaceae: Mimosoideae). Cytologia 66:
11-17.
Theplantlist www.theplantlist.org/tpl/search?q=
Aeschynomene
Verdcourt B. 1971. Aeschynomene. In: Gillet JB, Polhill
RM, Verdcourt B (Eds) Flora of Tropical East Africa,
Leguminosae, Papilionoideae. Royal Botanic Gar-
dens, Kew Publishing, London, 364-406.
Wojciechowski MF, Lavin M, Sanderson MJ. 2004. A
phylogeny of legumes (Leguminosae) based on analy-
sis of the plastid matK gene resolves many well-sup-
ported subclades within the family. American Journal
of Botany 91: 1846-1862.
Xiong Z, Pires JC. 2011. Karyotype and identication
of all homoeologous chromosomes of allopolyploid
Brassica napus and its diploid progenitors. Genetics
187: 37-49.