Available via license: CC BY
Content may be subject to copyright.
J.Fungi2020,6,101;doi:10.3390/jof6030101www.mdpi.com/journal/jof
Review
MolecularMethodsfortheDiagnosis
ofInvasiveCandidiasis
IrisCamp,KathrinSpettelandBirgitWillinger*
DivisionofClinicalMicrobiology,DepartmentofLaboratoryMedicine,MedicalUniversityofVienna,
1090Vienna,Austria;iris.camp@meduniwien.ac.at(I.C.);kathrin.spettel@meduniwien.ac.at(K.S.)
*Correspondence:birgit.willinger@meduniwien.ac.at
Received:25May2020;Accepted:4July2020;Published:6July2020
Abstract:InvasiveinfectionscausedbymembersofthegenusCandidaareontherise.Especially
patientsinintensivecareunits,immunocompromisedpatients,andthoserecoveringfrom
abdominalsurgeryareatriskforthedevelopmentofcandidemiaordeep‐seatedcandidiasis.Rapid
initiationofappropriateantifungaltherapycanincreasesurvivalratessignificantly.Inthepast,
mostoftheseinfectionswerecausedbyC.albicans,aspeciesthattypicallyisverysusceptibleto
antifungals.However,inrecentyearsashifttowardsinfectionscausedbynon‐albicansspecies
displayingvarioussusceptiblypatternshasbeenobservedandthepromptdiagnosisofthe
underlyingspecieshasbecomeanessentialfactordeterminingthetherapeuticoutcome.Thegold
standardfordiagnosinginvasivecandidiasisisbloodculture,eventhoughitssensitivityislowand
thetimerequiredforspeciesidentificationusuallyexceeds48h.Toovercometheseissues,blood
culturecanbecombinedwithothermethods,andalargenumberoftestshavebeendevelopedfor
thispurpose.Theaimofthisreviewwastogiveanoverviewonstrengthsandlimitationsof
currentlyavailablemolecularmethodsforthediagnosisofinvasivecandidiasis.
Keywords:Candida;invasive;diagnosis;molecular;candidemia;T2
1.Introduction
ThegenusCandidacomprisesadiversegroupofdimorphicfungithatarecommensalinhabitants
ofmucousmembranes[1]somespecies,likeC.parapsilosis,additionallycanbefoundascolonizers
onthehumanskin[2].Candidaspecies,therefore,areoftenisolatedfromnon‐sterileclinicalsamples,
suchasswabsfromthegastrointestinalorurogenitaltract.Eventhoughthesefindingsusuallydo
notholdanypathologicvalueinasymptomaticimmunocompetentpatients,Candidacancause
invasiveinfectionsthataremarkedbyhighmortalityrates[3,4].Especiallypatientsonintensivecare
units(ICU),aswellasimmunosuppressedorneutropenicpatients,areathigherriskforthe
developmentofaninvasivecandidiasis(IC)[5],andimprovedtreatmentstrategiesandsurvivalrates
foranumberofseverediseaseslikehematologicmalignancieshaveledtoanevergrowingpoolof
patientssusceptibletoandaffectedbyinvasivefungaldiseases[6].Thepopulation‐basedincidence
ofIChasbeenreportedtobebetween1.9and24cases/100,000/peryear[7–16].Worldwide,over
250,000casesofICandmorethan50,000deathsperyearareduetotheseinfections[17].ICoften,but
notalways,goesalongwithcandidemia,andindeedCandidaspp.havebeenreferredtobeingthe
fourthmostcommoncauseofnosocomialbloodstreaminfections[18].
Inmostcases,ICoriginatesfromthepatient’sownflora,andtheriskforthedevelopmentof
suchaninfectionincreaseswiththenumberofbodysitescolonizedbyCandida[19–21].Eventhough
C.albicansisstillthespeciesresponsibleformostcasesofIC,moreandmoreinvasiveinfectionsdue
tonon‐albicansspecieshavebeennotedinrecentyears[22–27].Thisisrelevantasnotonlyvirulence
andpathogenicitybutalsoresistanceprofilesvarybetweenspecies.WhileC.albicansusuallyis
susceptibletoallmajorgroupsofantifungals,C.glabratacanacquireresistancetoazoles[28],C.
J.Fungi2020,6,1012of14
parapsilosisandC.guilliermondiitoechinocandins[29,30],C.lusitaniaemaybelesssusceptibleto
amphotericinB[31],andC.kruseiisolatesareintrinsicallyresistanttofluconazole.Thus,theobserved
shifttowardsnon‐albicansspeciesmakesitmoredifficulttochoosetheappropriateempirictherapy.
AnotherconcernistheemergenceofC.auris.Thisoftenmulti‐resistantpathogenwasfirstdescribed
inJapanin2009[32].Sincethen,severaloutbreakshavebeenreported[33–35].
ThespectrumofclinicalsignsandsymptomsofICiswideandcanbeunspecific,butaninvasive
fungaldiseaseshouldalwaysbeconsideredifthepatient’sconditiondoesnotimproveunder
antibiotictherapy,especiallyifcolonizationwithCandidaspp.hasbeenobservedinahigh‐risk
patient.ToimprovetheoutcomeofIC,thepromptinitiationofanappropriateantimycotictherapy
isessential[36].Giventhedifferencesinresistancepatterns,fastspecies‐levelidentificationis
requiredforchoosingthecorrectantimycoticagentwhenresultsofantimicrobialsusceptibility
testingarenotyetavailable.
Cultureremainsoneofthekeymethodsfordiagnosingafungalinfection.However,the
definitivetreatmentofICisoftendelayedbytheinsensitivityofculture,andthisdelaymayleadto
highmortalityrates(35–75%)[37].Eventhoughbloodcultures(BC)aresensitiveatdetectingviable
Candidacells,withalimitofdetectionofonecolonyformingunit(CFU)/mL,theiroverallsensitivity
acrossthespectrumofICisonly50%,andtheyhavealagtimeforidentificationofupto5days
[38,39].Nevertheless,BCiscurrentlyconsideredthe“goldstandard”intheeventofanysuspected
caseofinvasivefungalinfection,butthecombinationofculturewithothermethodscanfacilitatea
timelierdiagnosis.Molecularamplificationtechniquesenablefastandsensitivedetectionand
identificationbydirectlydetectingandanalyzingtinyamountsoffungalDNApresentinaclinical
samplewithouttheneedforpriorcultivation,whichmakesthesetestsappealingfortheearly
diagnosisofIC,particularlyforcasesofICthataremissedbyculture.MultiplePCRassaystargeting
variousgeneticsequences(18SrDNA,28SrDNA,5.8SrDNA,internaltranscribedspacerregionsand
mitochondrialDNA)havebeendevelopedforthedetectionofabroadrangeoffungiindifferent
specimenssuchasblood,serum,plasma,bronchoalveolarlavage(BAL),sterilefluidsandtissues.
Dependingontheprimersused(i.e.primerstargetingeitherconservedorspecies‐specificregions),
fungalpathogenscanbedetectedinapanfungaloramorespecificmanner.Thesensitivityand
specificityofthevarioustechniquesarevariable,butmostlyanimprovedsensitivityisobserved
whencomparedtoclassicalculturalbasedmethods[40].Apartfromthedetectionandanalysisof
nucleicacids,molecularassayscanalsobebasedonproteomicprofiling.Inthisreview,wewillfocus
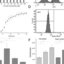
oncommerciallyavailabletests(Figure1).
Figure1.OverviewofavailablemoleculartestsforthediagnosisofinvasiveCandidainfections.BDG:
beta‐D‐glucan;
a
wholeblood,
b
wholeblood,plasmaandserum,
c
plasmaandsyntheticBAL,
d
various
clinicalsamples,
e
extractedDNA
J.Fungi2020,6,1013of14
Table1.Listofbloodculture‐dependenttests.
ProductManufacturerCandidaspp.
Detected
Assay
TimeMethodApproval
noGramstainrequired
SepsiTyper®Bruker
Daltonicspan‐Candida15–20
min
protein
extraction
followed
by
MALDI‐
TOFMS
CE/IVD
FilmArray®BCID
PanelBiomerieux
C.albicans,
C.glabrata,
C.parapsilosis,
C.tropicalis,
C.krusei
60minMultiplex
PCRCE/IVD
Accelerate
PhenoTestTMBCKit
Accelerate
Diagnostics
C.albicans,
C.glabrata90minautomated
FISHCE/IVD
SepsisFlowChipMaster
DiagnosticaC.albicans3hMultiplex
PCRCE/IVD
Gramstainrequired
CandidaQuickFISH®OpGen
C.albicans
C.parapsilosis,
C.glabrata
20minFISHCE/IVD
YeastTrafficLight
PNAFISH®OpGen
C.albicans/C.
parapsilosis,
C.tropicalis,
C.glabrata/C.
krusei
90minFISHCE/IVD
eplex®BCIDFPPanelGenMarkDx
C.albicans,
C.auris,
C.dubliniensis,
C.famata,
C.glabrata,
C.guilliermondii,
C.kefyr,
C.krusei,
C.lusitaniae,
C.parapsilosis,
C.tropicalis
90minMultiplex
PCRCE/IVD
MALDI‐TOFMS:matrix‐assistedlaserdesorption/ionizationtimeofflightmassspectrometry;
CE/IVD:ConformitèEuropëenne/invitrodiagnostic;FISH:fluorescenceinsituhybridization.
2.BloodCulture‐DependentMolecularDiagnostics
Thesetestsystems(Table1)aredesignedforusewithaliquotsofpositivebloodculturesamples.
Thus,thetimerequiredforapositivebloodculturecannotbeeliminated.Asthepathogenloadis
highinpositivebloodculturebottles,sensitivityisnotachallengefortheseassays.
TheMALDISepsityper®IVDKit(BrukerDaltonics,Bremen,Germany)allowsforpathogen
identificationviaMALDI‐TOFMSanalysisfromaliquotsofpositivebloodculturebottlesafterashort
proteinextraction.Inarecentlypublishedstudy[41],62.5%ofallCandidaisolatescouldbeidentified
withtheMALDISepsityper®IVDKitdirectlyfrompositivebloodculturebottles.IfaBruker
Biotyperinstrumentisavailable,thistestisaratherinexpensivealternativetotestsbasedonnucleic
aciddetection.Anotherbenefitisthepotentialtoidentifyallpathogensincludedinthedatabase;
thus,rareyeastsincludingC.auriscanalsobeidentified.
J.Fungi2020,6,1014of14
TheFilmArray®BCIDPanel(Biomerieux,Marcyl’Etoile,France)isaConformitèEuropëenne/in
vitrodiagnostic(CE/IVD)‐certifiednestedmultiplexPCRsystem.Twenty‐fourpathogens,including
thefivemostcommonCandidaspp(C.albicans,C.glabrata,C.parapsilosis,C.tropicalis,andC.krusei),
canbedetectedbytheassaywithminimalhands‐ontimeandaturn‐aroundtimeof1h.Inastudy
withbothclinicalandspikedsamples,asensitivityof99.2%andaspecificityof99.9%wereobserved
forCandidaspp.whenresultsoftheBCIDpanelwerecomparedwithconventionalculture[42].In
anotherrecentlypublishedstudy,theFilmArray®BCIDpanelwasperformedon85positiveblood
cultureswithyeastsvisibleintheGramstain.Atotalof91yeaststrainswereisolatedbyculture,and
84oftheisolatesbelongedtooneofthefiveCandidaspeciescontainedinthepanel.Allofthosewere
identified.Sevenisolatesbelongedtospeciesnottargetedbythetest;thosewerenotdetected.Ten
bloodculturescontainedmorethanonepathogen,andallpathogensincludedinthepanelwere
identifiedcorrectly[43].SincethetestpanelincludesyeastsaswellasGram‐positiveandGram‐
negativebacteria,itisnotnecessarytoperformaGram‐stainpriortotheassay.
TheAcceleratePhenoTestTMBCKit(AccelerateDiagnostics,Tuscon,AZ,USA)isanother
CE/IVDapprovedtestsystemthatallowstheidentificationandrapidphenotypicantimicrobial
susceptibilitytestingofseveralGram‐positiveandGram‐negativebacteriainpositivebloodcultures.
Inadditiontobacterialpathogens,thesystemcandetectC.albicansandC.glabrata.However,rapid
susceptibilitytestingisnotavailableforfungi.ThetestwasrecentlyevaluatedinastudybyBurnham
etal.[44];10ofthe125bloodculturespositiveforpathogensincludedinthetest’spanelcontained
C.glabrata,and5culturescontainedC.albicans.TheassayfailedtodetectC.glabrataintwoofthese
samples,whilethreefalsepositiveresultswithC.glabrata,aswellasonefalsepositiveC.albicans
result,werereported.Thus,forC.albicans,asensitivityof100%andaspecificityof99.3%were
reported,whilethesensitivityandspecificityforC.glabratawere80.0%and97.9%respectively.Five
falsepositiveresultsforC.glabratawerealsoreportedinastudypublishedin2017[45].Atthetime,
thisunfavorableoutcomewasexplainedbytheuseoftheoldersoftwareversion(v1.0),anditwas
discussedthatsoftwareupdatesmightresolvetheissue.
TheSepsisFlowChip(MasterDiagnostica,Granada,Spain)isaCEandIVDapprovedmultiplex
PCRtestwhichisabletodetect22resistancegenesinadditiontomorethan40pathogensincluding
C.albicansfrompositiveBCinthreehours.Thisisachievedbytheuseofbiotinylatedprimers,
automatedreversehybridizationtoachipmembraneandsubsequentimmunoenzymaticdetection
ofpositivesignals.Inthefirstevaluationonclinicalsamples[46],sixyeast‐positiveBCwereincluded.
FiveofthesecontainedC.albicans,andallweredetectedbythetest.OneBCcontainedC.parapsilosis;
forthissample,theassaydidnotyieldapositiveresult.Thus,forC.albicans,asensitivityof100%
wasreported.
TheCandidaQuickFISH®andtheYeastTrafficLightPNAFISH®(OpGen,Gaithersburg,MA,
USA)areCE/IVDcertifiedtestsusingpeptidenucleicacidprobesforfluorescenceinsitu
hybridization.Theycanbeperformeddirectlyonaliquotsfromyeastpositivebloodculturebottles
(i.e.whenyeastcellshavebeenobservedintheGramstain).WiththeYeastTrafficLightPNAFISH®,
identificationofC.albicans/C.parapsilosis,C.tropicalis,andC.glabrata/C.kruseiispossiblewithin90
min,whiletheCandidaQuickFISH®candetectC.albicans,C.parapsilosis,andC.glabratawithin20
min.Inalargeevaluationstudywith216bloodculturesamples[47],theYeastTrafficLightPNA
FISH®yieldedthecorrectresultin96%ofcases.OneisolateofC.parapsilosiswasmisidentifiedasC.
tropicalis,andafalsenegativeresultwasobtainedforonecaseofC.parapsilosisandonecaseofC.
tropicalis.Additionally,crossreactivitywithC.bracarensis,C.nivariensis,C.orthopsilosis,N.delphensis,
andR.mucilaginosawasobserved.Drawbacksmightincludethelimitedspectrum,aswellasthe
requirementforafluorescencemicroscope.
FortheePlex®BCID(GenMarkDX,Carlsbad,CA,USA),arecentlyCE/IVDcertifiedsystem,
thereisachoiceofthreepanels.Thus,dependingontheresultsoftheGram‐stainperformedon
positivebloodcultures,therespectivepanelwillbeused.Thefungalpathogen(FP)paneltargets11
Candidaspp.(C.albicans,C.auris,C.dubliniensis,C.famata,C.glabrata,C.guilliermondii,C.kefyr,C.
krusei,C.lusitaniae,C.parapsilosis,andC.tropicalis)aswellasCryptococcusgattii,Cryptococcus
neoformans,FusariumandRhodotorula,andprovidesresultsin1.5h.Huangetal.tested210positive
J.Fungi2020,6,1015of14
bloodculturesfrompatientswithbloodstreaminfectionswiththeappropriatepanel.Yeastswere
onlyobservedintheGramstainofsevensamples;sixofthesesamplescontainedCandidaspp.
includedinthepanel,andallofthosewereidentifiedbythetest.OnesamplecontainedC.
inconspicua,whichisnotincludedinthepanelandthuswasnotidentified[48].
Table2.Listofbloodcultureindependenttests.
ProductManufacture
r
Candidaspp.
Detected
Assay
TimeMethodApproval
DNAextractionsteprequired
Singletarget:
AurisID®Olm
DiagnosticsC.auris45minaqPCRCE/IVD
Fungiplex®
Candidaauris
Bruker
DaltonicsC.auris<2haReal‐time
PCRRUO
Multiplextests:
CandID®Olm
Diagnostics
C.albicans
C.glabrata
C.parapsilosis
C.krusei
C.dubliniensis
C.tropicalis
45minaMultiplex
qPCRCE/IVD
Fungiplex®
Candida
Bruker
Daltonics
C.krusei
C.glabrata
Candidaspp.
(including:C.
albicans,
C.parapsilosis,
C.tropicalis,
C.dubliniensis)
<2ha
Multiplex
real‐time
PCR
CE/IVD
Fungiplex®
Universal
Bruker
DaltonicsCandidaspp.<2ha
Multiplex
real‐time
PCR
RUO
MycoRealCandida Ingenetix
C.albicans,
C.dubliniensis,
C.glabrata,
C.krusei,
C.lusitaniae,
C.parapsilosis,
C.tropicalis
2ha Multiplex
PCR RUO
MagicplexTM
SepsisSeegene
C.albicans
C.tropicalis
C.parapsilosis
C.glabrata
C.krusei
6hbMultiplex
PCRCE/IVD
Broad‐spectrumtests:
HybcellPathogens
DNAxBCubeDx
Microarray:
C.albicans
C.dubliniensis
C.parapsilosis
C.tropicalis
C.glabrata
+1panfungal
target
6hc
Panfungal
PCR(28s)&
Microarray
CE/IVD
J.Fungi2020,6,1016of14
Sequencing:
pan‐Candida
SepsiTestTM‐UMD
Molzym
Molecular
Diagnostics
pan‐Candida24h
Broad‐
spectrum
PCR(18S)
CE/IVD
MycoRealFungiIngenetixpan‐Candida24h
Broad‐
spectrum
PCR(ITS2)
RUO
Fullyautomated:
T2CandidaPanelT2
Biosystems
C.albicans/tropicalis
C.glabrata/krusei
C.parapsilosis
3–5h
Multiplex
PCR
followedby
automated
T2MRbased
detection
CE/IVD
aexcludingDNAextraction;bincludingDNAextraction;cifsequencingisnotrequired.CE/IVD:
ConformitèEuropëenne/invitrodiagnostic;RUO:researchuseonly;ITS:internaltranscribedspacer
2region.
3.BloodCultureIndependentMolecularDiagnostics
Bloodcultureindependentmolecularassays(Table2)canbeperformeddirectlyonwholeblood,
serum,orplasmasampleswithouttheneedtowaitforpositivebloodcultures.Thus,thetimesaving
potentialishigherthanwithbloodculturedependenttestsystems.Sincethepathogenloadinthe
bloodislow,thesensitivityofthesetestsystemscanbeanissue.Alargenumberofassaysisavailable
todayforthemoleculardiagnosisofIC;however,manyarein‐houseassaysorcommercially
availableresearch‐use‐onlytests.Broadspectrumtests,aswellastargetedmultiplexassays,are
available.Theprincipleofabroadspectrum/panfungaltestistoamplifyconservedtargetregions
thattheoreticallycanbefoundacrossallfungalspecies.Forspeciesidentification,obtainedamplicons
havetobeanalyzedfurther.Panfungalassaysgenerallyarelesssensitivethanassaystargeting
certainpathogenswithspecies‐specificprimers,buthavetheabilitytodetectallfungalpathogens,
notjustthemostfrequentones.AnissuewiththeuseofmultiplexPCRcanarisefromthefactthat
cliniciansarenotusuallyfamiliarwiththetestpanelsandthuscouldassumethatanegative
multiplexresultissufficientforrulingoutaninvasivefungalinfection.Therefore,itisimportantto
specifywhichpathogensarecoveredonthereportscreatedbytheclinicalmicrobiology/mycology
laboratory.
SomeofthetestsdescribedherecomewiththeirownDNAextractionkit,whileanumberof
differentextractionkits/protocolsarerecommendedforothertests.Asfungalcellsaredifficultto
lyse,theprotocolusedfortheextractionofnucleicacidsmighthavealargeeffectontheassay’s
outcome.Therefore,assayslackingtheirownDNAextractionmethodcanbemoredifficultto
standardize.
CandID®undAurisID®(OlmDiagnostics,NewcastleuponTyne,England)aretwonew
CE/IVDcertifiedqPCRteststhatcanbeperformedwithvariousreal‐timePCRinstruments.The
CandIDkitdetectsC.albicans,C.glabrata,C.parapsilosis,C.krusei,C.dubliniensis,andC.tropicalis;the
AurisID®kitdetectsC.aurisonly.Resultsareavailablewithin45minfromnucleicacidextraction;
noextractionprotocol/kitsarerecommended.Accordingtothemanufacturer,bothkitshavebeen
validatedwithfungalcultures,theCandID®kithasadditionallybeenvalidatedwithplasmaand
syntheticBALsamples,theAurisID®kitwithbloodsamples.Tothebestofourknowledge,no
studiesevaluatingtheclinicalperformanceoftheassayshavebeenpublishedsofar.
TheFungiplex®CandidaIVDPCRKit(BrukerDaltonik,Bremen,Germany)detectsC.krusei,C.
glabrataandCandidaspp.(including:C.albicans,C.parapsilosis,C.tropicalis,andC.dubliniensis)in
wholeblood,plasmaandserum.ForDNAextraction,kitsfromQiagenandBiomerieuxare
recommended,andtheassaymanualprovidesinstrumentsettingsforanumberofdifferent
thermocyclers.InasmallprospectivestudyonICUpatientswithsuspectedIC,theFungiplex®
J.Fungi2020,6,1017of14
CandidadetectedeightoutofeightpatientswithICandreachedasensitivityof100%anda
specificityof94.1%[49].BrukeralsooffersthepanfungalFungiplex®UniversalRUOPCRKitand
theFungiplex®CandidaAurisRUOPCRKitforusewithextractedDNA.
TheMagicplexTMSepsisReal‐timeTest(Seegene,Seoul,SouthKorea)isaCE/IVDapproved
multiplexrealtimePCRdetecting90pathogensatgenuslevel,and27pathogens,includingfive
Candidaspp.(C.albicans,C.tropicalis,C.parapsilosis,C.glabrata,C.krusei),atspecieslevelwithinsix
hoursfromwholeblood.TheSeegeneBloodPathogenKitTMisusedforthepre‐treatmentand
extractionofDNA,andthisstepisfollowedbyaconventionalPCR(onetubeforGram‐positive
bacteriaandresistancemarkersandonetubeforGram‐negativebacteriaandfungi)foramplicon
generation.Ifampliconsaredetected,theconventionalPCRisfollowedbytworeal‐timePCRsfor
screeningandspecieslevelidentification.Seegeneofferssoftware(SeegeneViewer)forthe
interpretationofresults.DeninaetalcomparedtheMagicplexTMtesttobloodculturein150samples
from89patients.Candidaspp.weredetectedbytheMagicplexTMinfoursamples;onlyoneofthese
sampleswasaccompaniedbyapositivebloodculture[50].Inarecentlypublishedstudy,14patients
withICwereincluded.Innineofthesepatients,Candidawasonlydetectedbybloodculture,intwo
patientsonlybytheMagicplexTMassay,andinthreepatientsbybothmethods[51].Importantly,the
twoisolatesdetectedonlybytheMagiplexTMassaybelongedtoC.parapsilosis,whichiswellknown
tobeacolonizingspecies.Thus,thedetectionofC.parapsilosishastobeinterpretedwithcaution.
Moreover,theauthorsdescribethatthetest’slowsensitivitymakesitsimplementationasaroutine
testinclinicalmicrobiologylaboratoriesdifficult.
TheMycoRealCandida(Ingenetix,Vienna,Austria)isaresearch‐use‐onlymultiplexPCRfor
thedetectionofC.albicans,C.dubliniensis,C.glabrata,C.krusei,C.lusitaniae,C.parapsilosis,andC.
tropicalis.Inthisassay,species‐specificbiprobesareused.Thetestwasevaluatedinastudyusing
bothspikedandclinicalsamples[52].Resultsoftheanalyticalandclinicalevaluationshowedthat
thisassaywashighlysensitiveandcanbeusedinclinicallaboratoriesasasimplescreeningtestfor
thementionedCandidaspecies.IngenetixalsoofferstheMycoRealFungi,aresearch‐use‐only
panfungaltesttargetingtheinternaltranscribedspacer(ITS)2region.Theassaykitcontainsprimers,
probes,andapositivecontrol,whilethereactionmixhastobeprovidedbytheuser.ForDNA
extractionfromsamples(blood,sterilefluids,tissue,paraffinembeddedtissue,andBAL),amodified
protocolforusewiththeHighPurePCRTemplatePreparationKitfromRocheDiagnosticsis
recommended.ThesystemhasbeenvalidatedfortheLightCycler®2.0instrument(Roche
Diagnostics),andaLoD95%of15CFU/PCRisreportedbythemanufacturer.Obtainedamplicons
havetobesequencedforspeciesidentification.Thistestwasbasedonanin‐housetest[53,54].
TheSepsiTestTM–UMD(MolzymMolecularDiagnostics,Bremen,Germany)isasystemforthe
CE/IVDcertifiedbroad‐rangedetectionofintactbacterialandfungalpathogenswithananalytical
sensitivityrangingfrom10to80CFU/mL.Inadditiontowholebloodsamples,thistestisvalidated
forsterilefluids,tissuesamples,andswabs.ForpathogenenrichmentandDNAextraction,Molzym
offersanautomatedsolution,inwhichthesestepsareperformed,fullyautomatedbytheSelectNATM
plusrobot(Micro‐DxTMCEIVD).Asemi‐automatedsolution(UMD‐SelectNATMCEIVD)with
manualpathogenenrichment,followedbyautomatedDNAisolationononeofthefollowing
instruments—Liaison®Ixt(Diasorin),Arrow®(Nordiag),Seeprep12TM(Seegene),orGenoXtract®
(HainLifescience)—aswellasmanualextractionarepossible.Afterwards,16Sand18SrRNAgenes
areamplifiedintwoseparatereactions.Obtainedampliconshavetobesequenced,andsequences
arethenanalyzedwiththefreeonlineSepsiTestTM‐BLASTtool.Themajoradvantageofthisbroad‐
rangetestisitswidespectrumwhichincludesfastidiousorganismsthatarenotdetectablebyculture.
IncaseofapositivePCRresult,theneedforsequencingincreasesthetimetogaintheresult.Even
thoughseveralstudieshaveevaluatedtheperformanceofthetest[55–59],fewcasesofICwere
includedinthesestudies.Schreiberetal.reportedonecaseofC.albicansdetectedinbloodcultures
ofapatientwithanegativePCRresult[56],andNiemanetal.foundC.albicansintwopatients,the
yeastwasonlydetectedbybloodculturesinone,andonlyinthePCRassayinthesecondpatient
[58].
J.Fungi2020,6,1018of14
TheHybcellPathogensDNAxB(CubeDx,St.Valentin,Austria)isarecentlyCE/IVDapproved
testforthedetectionofbacteria,resistancegenes,andfungi.DNAisisolatedfrom500mLwhole
bloodwiththeGINApathogenenrichmentandDNApurificationkit(CubeDx).Subsequently,four
separatePCRreactionsarecarriedout(positivecontrol,bacterialpanel:16SrDNA,fungalpanel:28s
rDNA,andpanelfortheresistancemarkersvanA,vanB,mecA,andmecC)andafluorescentdyeis
incorporatedintoampliconsduringthePCR.UponcompletionofthePCR,PCRproductsare
transferredtocylindricalmicroarrays—so‐calledhybcells—andampliconsareidentifiedinthe
hyborgdevicebybindingtoimmobilizedprobesviaelongationanddetectionoffluorescencesignals.
Thesystemcanidentify1panbacterialtarget,4bacterialgeneraand28bacterialspecies,aswellas1
panfungaltarget,2fungalgenera,and13fungalspecies.Thus,sequencingofthePCRproductsisnot
necessaryifthepathogenisincludedinthetestpanel,andresultscanbeavailablewithin3h.Should
specieslevelidentificationyieldnoresultinasamplepositiveforthepanbacterialorthepanfungal
target,leftoverPCRproductscanbesubjectedtoSangersequencing.Thistestiscurrentlyunder
evaluation.Sofar,nopeer‐reviewedstudyresultsareavailable.
TheT2CandidaPanel(T2Biosystems)isaCE/IVDapprovedtestforuseontheT2Dxinstrument,
whichutilizesT2MagneticResonance(T2MR).TheCandidapanelcandetectthreegroupsofCandida
(C.albicans/C.tropicalis,C.glabrata/C.krusei(whichalsoincludesS.cerevisiaeandC.bracarensis),and
C.parapsilosis(whichincludesC.orthopsilosisandC.metapsilosis))inEDTAbloodsampleswith
minimalhands‐on‐time.Afterin‐cartridgeDNAextraction,theITS2regionisamplified.Amplicons
aredetectedviahybridizationwithspecificcaptureprobescarryingsuperparamagneticparticles.The
resultingagglomerationoftheseparticlesinducesashiftinthesample’smagneticresonance.This
methodisabletodetectminimalamountsofintacttargetcells(1CFU/mL)—butnotfreeDNA—with
atimetoresultof3–5h.Neelyetal.[60]evaluatedthemethodforthedetectionofCandidain2013.
Intheirstudy,wholebloodsampleswerespikedwithdifferentconcentrationsofCandidaspp.
includedinthepanel;highagreementratesbetweentheT2MRandBC(97.8%positiveand100%
negativeagreement)wereobserved.Mylonakisetal.laterevaluatedthetestinalargeprospective
studywithsamplesfrom1801patients;250ofthesesampleswerespikedwithCandidaspp.[61].The
overallanalyticalsensitivityoftheT2MRwas91.6%anditsspecificitywas99.4%.In31cases,T2MR
andBCdidnotyieldthesameresults;2patientswerepositiveinBCbutnotwiththeT2MR,while
samplesfrom29patientswerepositivewithT2MRbutnegativeinBC.Arendrupetal.recently
conductedanotherprospectivestudywith126ICUpatientswhichwereclassifiedintogroupsof
proven,likely,possible,orunlikelyICbasedontheresultsofBC,culturefromsterilesites,
colonization,aCandidaantigenassay,andclinicalfindings.ComparedwithBCandtheantigentest,
theT2CandidaPanelhadthehighestsensitivityforthedetectionofIC[62],eventhoughthe
sensitivitywaslowerthanobservedinthe2015Mylonakisetal.study[61].Thebestsensitivitywas
achievedbyacombinationofBCandtheT2CandidaPanel[62].Asthetestalsodetectsnon‐viable
cells,T2MRmightbeausefultoolforthemonitoringofcandidemiauponinitiationofantifungal
therapy.ThiswasdemonstratedinastudypublishedbyMylonakisetalin2018[63].Follow‐ up
bloodsamples(bloodculturesandwholeblood)from31patientswithcandidemiawereincluded.
ThirteenpatientshadatleastonepositiveT2MRresult,whileBConlydetectedthepresenceof
Candidain4ofthese13patients.Clancyetal.[64]comparedthepositivityoffollow‐upsamplesfrom
152patientswithcandidemia.Duringasecondblooddraw,samplesforT2MRandacompanionBC
(cBC)wereobtained.SamplesfortheT2werefrozenandanalyzedinbatches.Inpatientsunder
antifungaltherapy,theT2MRassaywasmoreoftenpositivethanthecBC(50%vs.21%),whereasno
differencewasobservedinuntreatedpatients.Invalidreportswerereportedin9%ofT2samples;
thismightbeduetotheanalysisoffrozensamples.Thetestonlydetectsintactorganisms(notfree
DNA),andthereforeshouldnotbeperformedonfrozensamples.Thus,studiesworkingwithfrozen
samplesmightnotreflecttrueperformancecharacteristics.Zurletal.analyzedfrozensamplesfrom
32patientswithcandidemiaandfrom22patientswithdeep‐ seatedcandidiasis[65].Samplesfor
T2MRtestingwerecollectedatvarioustimepointsrangingfrom2daysbeforeuntil5daysafterthe
indexculture(CandidapositiveBCorsterilesiteculture).Severalinvalidsamples/instrumenterrors
wereobserved.Furthermore,eightsampleswhichwerecollectedconcurrentlywiththepositive
J.Fungi2020,6,1019of14
indexBCyieldednegativeT2MRresults.Intwocases,theT2detectedaCandidaspeciesdifferent
fromthespeciesfoundintheBC.Inthegroupofpatientswithdeep‐seatedcandidiasis,the
T2Candidapanelgaveatleastonepositiveresultinsixpatients(27.3%).Remarkably,allBCs
collectedfrompatientswithdeep‐seatedcandidiasisremainednegative.Thus,eventhoughthe
percentageofpositiveT2resultsdoesnotseemveryhigh,thisisaninterestingobservation,sincethe
diagnosisofdeep‐seatedcandidiasisisverychallenging.InadditiontotheT2CandidaPanel,
T2Biosystemsalsooffersaresearch‐use‐onlypanelforthedetectionofC.auris[66]inskinswab
samples.InadditiontothediagnosticperformancetheroleofT2Candidaasaprognosticandpatient
managementtoolshouldalsobeevaluated.Munozetal.showed,inaprospectiveobservational
multicenterstudyofpatientsreceivingdefinitiveantifungaltreatmentforcandidemia,thatapositive
T2MRwasassociatedwithahigherriskofpooroutcome,whilethedetectionofbeta‐D‐Glucandid
notcorrelatewiththeoutcome.ApositiveT2Candidaresultwithinthefirst5daysafterthereportof
apositiveBCwasanindependentriskfactorforcomplicatedcandidemia,definedbyattributable
mortalityordevelopmentofmetastatic,deep‐seatedinfection[67].Inanothermulti‐center
investigation,Munozetal.showedthatT2Candidaperformedinpatientswithprovencandidemia
maybeabettermarkerofcomplicatedinfectionthanfollow‐upbloodculturesordetectionofbeta‐
D‐glucan(BDG).TheseresultsindicatethatT2Candidamayinfluencethelengthandtypeof
antifungaltherapyinthispopulationandmightbeusedinthesenseofantimicrobialstewardship
[68].Asaconsequence,testresultscouldbeusedtoexpediteantifungaltreatmentofcandidemia,
andreduceoverallantifungalusagewithoutanegativeeffectonpatientoutcomes.UsingT2Candida
incombinationwithculturesislikelytooffergreatestvalue.However,antifungaltherapymay
influencethesensitivityoftheperformanceoftheT2CandidaPanel.AsdescribedbyClancyetal.
T2Candidashowedlimitedsensitivity(36%)/negativepredictivevalue(NPV)(80%)intheMADRID
prospectiveobservationalstudyundertheinfluenceofempiricalantifungaltherapy,whereas
specificity/positivepredictivevalue(PPV)wasexcellent(100%),indicatingarolebettersuitedto
confirmingadiagnosisorpersistentinfection[69].Ashasbeennoted,stratificationofhigh‐risk
patientsthroughrisk‐predictionmodelingisessentialtoachieveasufficientpre‐testprobability.
Irrespectiveoftheprevalenceofdisease,theNPVoftheT2testis>98%,butaprevalenceofaround
10%maybeoptimal,providingaPPVandNPVofapproximately82%and99%,respectively[69,70].
4.Summary
Diagnosingfungalinfectionshasalwaysbeenchallenging.Advancesinmoleculardiagnostic
technologieshavegeneratedarangeoftestswithrapidturnaroundtimesforthediagnosisand/or
screeningofpatientsatriskforinvasivefungalinfections.IncreasingexperiencewithPCRassaysfor
thedirectdetectionoffungiinclinicalspecimensandavailableclinicalvalidationstudieshave
positionedtheseassaytypeswellonthewaytobecomingroutineinclinicallaboratories.
SeveralPCRassays,includingcommerciallyavailablekits,havebeendevelopedforthe
detectionofCandidaspp.inpatientswithcandidemiaandIC.Thehighsensitivitymakestheseassays
appealingtoolsfortheearlydiagnosisofIC.DependingonthemethodusedforDNAextraction,free
DNAorintactpathogencellsaredetected.Thisdifferencecanberelevantforpatientsunder
antifungaltherapyasthismayspecificallyinfluencetheoutcomeofmolecularteststhatdetectonly
intactcells.Ontheotherhand,theinterpretationofpositiveresultsfromassaysdetectingfreeDNA
canbechallenging.Thus,thepositionofCandidaPCRassaysinthediagnosticalgorithmofICisnot
easytoestablish.Aspublisheddatashow,CandidaPCRhasahighersensitivitythanbloodculture
butshowsthebestefficacywhenusedinconjunctionwithbloodculturesand/oradditionaltestssuch
asthedetectionofBDG.Inaddition,theuseofmoleculartechniquesforpositivebloodcultures
allowsamorerapididentificationofCandidaspp.
Asearlyinitiationofeffectiveantifungaltherapyisassociatedwithimprovedoutcomes[71],it
iscrucialtostartatargetedtherapyasearlyaspossible.Directmoleculardetectionorrapid
identificationofCandidaspp.frombloodculturesbyuseofmolecularassaysshowsthepotentialfor
earlyadministrationofanoptimalantifungaltherapy.Inaddition,theseassaysmayallowforthe
J.Fungi2020,6,10110of14
correctchoiceoflengthandtypeofantifungaltherapy,andmaythusbeusedinthesenseof
antimicrobialstewardship.
However,manyoftheseassaysremainunderinvestigationastheyhavenotbeenvalidatedfor
diagnosingICinmulti‐centerstudies.Thechoiceofadoptinganin‐houseratherthancommercial
assayisdependentuponcosts,aswellasworkflowandcapacityinindividuallaboratories[72],and
resultsofmolecularassaysshouldalwaysbeinterpretedwithcaution[73].Therefore,moredatafrom
multi‐centerstudiesisneededforafinalassessmentofcommercialassays.
Authorcontributions:I.C.wrotethefirstdraftandadaptedthemanuscriptfollowingreviews,K.S.was
involvedinthereviewofexistingliterature,designedthefigureandtables,andreviewedthemanuscript,and
B.W.wasinvolvedinconceptualizing,writingandreviewingthemanuscriptbeforesubmission.Allauthors
havereadandagreedtothepublishedversionofthemanuscript.
ConflictsofInterest:TheauthorsarecurrentlyconductingastudyontheclinicalperformanceoftheT2Candida
Panel.Forthestudyperiod,theT2DxinstrumentandnecessarykitsaresuppliedbyT2Biosystems/Biomedica.
References
1. Bougnoux,M.‐E.;Diogo,D.;François,N.;Sendid,B.;Veirmeire,S.;Colombel,J.F.;Bouchier,C.;Van
Kruiningen,H.;d’Enfert,C.;Poulain,D.Multilocussequencetypingrevealsintrafamilialtransmissionand
microevolutionsofCandidaalbicansisolatesfromthehumandigestivetract.J.Clin.Microbiol.2006,44,
1810–1820.
2. Kühbacher,A.;Burger‐Kentischer,A.;Rupp,S.InteractionofCandidaSpecieswiththeSkin.
Microorganisms2017,5,32.
3. Morgan,J.;Meltzer,M.I.;Plikaytis,B.D.;Sofair,A.N.;Huie‐White,S.;Wilcox,S.;Harrison,L.H.;Seaberg,
E.C.;Hajjeh,R.A.;Teutsch,S.M.Excessmortality,hospitalstay,andcostduetocandidemia:acase‐control
studyusingdatafrompopulation‐basedcandidemiasurveillance.InfectControl.Hosp.Epidemiol.2005,26,
540–547.
4. Gudlaugsson,O.;Gillespie,S.;Lee,K.;Berg,J.V.;Hu,J.;Messer,S.;Herwaldt,L.;Pfaller,M.;Diekema,D.
AttributableMortalityofNosocomialCandidemia,Revisited.Clin.Infect.Dis.2003,37,1172–1177.
5. Bassetti,M.;Giacobbe,D.R.;Vena,A.;Trucchi,C.;Ansaldi,F.;Antonelli,M.;Adamkova,V.;Alicino,C.;
Almyroudi,M.‐P.;Atchade,E.;etal.Incidenceandoutcomeofinvasivecandidiasisinintensivecareunits
(ICUs)inEurope:resultsoftheEUCANDICUproject.Crit.Care2019,23,219.
6. Martin,G.S.;Mannino,D.M.;Eaton,S.;Moss,M.TheepidemiologyofsepsisintheUnitedStatesfrom1979
through2000.N.Engl.J.Med.2003,348,1546–1554.
7. Almirante,B.;Rodríguez,D.;Park,B.J.;Cuenca‐Estrella,M.;Planes,A.M.;Almela,M.;Mensa,J.;Sanchez,
F.;Ayats,J.;Gimenez,M.;etal.EpidemiologyandpredictorsofmortalityincasesofCandidabloodstream
infection:resultsfrompopulation‐basedsurveillance,barcelona,Spain,from2002to2003.J.Clin.Microbiol.
2005,43,1829–1835.
8. Arendrup,M.C.;Fuursted,K.;Gahrn‐Hansen,B.;Jensen,I.M.;Knudsen,J.D.;Lundgren,B.;Schønheyder,
H.C.;Tvede,M.SeminationalSurveillanceofFungemiainDenmark:NotablyHighRatesofFungemiaand
NumbersofIsolateswithReducedAzoleSusceptibility.J.Clin.Microbiol.2005,43,4434–4440.
9. Ásmundsdóttir,L.R.;Erlendsdóttir,H.;Gottfredsson,M.IncreasingIncidenceofCandidemia:Resultsfrom
a20‐YearNationwideStudyinIceland.J.Clin.Microbiol.2002,40,3489–3492.
10. Diekema,D.J.;Messer,S.A.;Brueggemann,A.B.;Coffman,S.L.;Doern,G.V.;Herwaldt,L.A.;Pfaller,M.A.
EpidemiologyofCandidemia:3‐YearResultsfromtheEmergingInfectionsandtheEpidemiologyofIowa
OrganismsStudy.J.Clin.Microbiol.2002,40,1298–1302.
11. Hajjeh,R.A.;Sofair,A.N.;Harrison,L.H.;Lyon,G.M.;Arthington‐Skaggs,B.A.;Mirza,S.A.;Phelan,M.;
Morgan,J.;Lee‐Yang,W.;Ciblak,M.A.;etal.IncidenceofbloodstreaminfectionsduetoCandidaspecies
andinvitrosusceptibilitiesofisolatescollectedfrom1998to2000inapopulation‐basedactivesurveillance
program.J.Clin.Microbiol.2004,42,1519–1527.
12. Kao,A.S.;Brandt,M.E.;Pruitt,W.R.;Conn,L.A.;Perkins,B.A.;Stephens,D.S.;Baughman,W.S.;Reingold,
A.L.;Rothrock,G.A.;Pfaller,M.A.;etal.TheepidemiologyofcandidemiaintwoUnitedStatescities:results
ofapopulation‐basedactivesurveillance.Clin.Infect.Dis.1999,29,1164–1170.
13. Laupland,K.B.;Gregson,D.B.;Church,D.L.;Ross,T.;Elsayed,S.InvasiveCandidaspeciesinfections:a5
yearpopulation‐basedassessment.J.Antimicrob.Chemother.2005,56,532–537.
J.Fungi2020,6,10111of14
14. Poikonen,E.;Lyytikäinen,O.;Anttila,V.‐J.;Ruutu,P.CandidemiainFinland,1995–1999.Emerg.InfectDis.
2003,9,985–990.
15. Sandven,P.;Bevanger,L.;Digranes,A.;Haukland,H.H.;Mannsåker,T.;Gaustad,P.Candidemiain
Norway(1991to2003):ResultsfromaNationwideStudy.J.Clin.Microbiol.2006,44,1977–1981.
16. Toda,M.;Williams,S.R.;Berkow,E.L.;Farley,M.M.;Harrison,L.H.;Bonner,L.;Marceaux,K.M.;Hollick,
R.;Zhang,A.Y.;Schaffner,W.;etal.Population‐BasedActiveSurveillanceforCulture‐Confirmed
Candidemia—FourSites,UnitedStates,2012–2016.MMWRSurveill.Summ.2019,68,1–15.
17. Kullberg,B.J.;Arendrup,M.C.InvasiveCandidiasis.N.Engl.J.Med.2016,374,794–795.
18. Wisplinghoff,H.;Bischoff,T.;Tallent,S.M.;Seifert,H.;Wenzel,R.P.;Edmond,M.B.Nosocomial
BloodstreamInfectionsinUSHospitals:Analysisof24,179CasesfromaProspectiveNationwide
SurveillanceStudy.Clin.Infect.Dis.2004,39,309–317.
19. Jordà‐Marcos,R.;Álvarez‐Lerma,F.;Jurado,M.;Palomar,M.;Nolla‐Salas,J.;León,M.A.;León,C.Risk
factorsforcandidaemiaincriticallyillpatients:aprospectivesurveillancestudy*.Mycoses2007,50,302–
310.
20. Pittet,D.;Monod,M.;Suter,P.M.;Frenk,E.;Auckenthaler,R.Candidacolonizationandsubsequent
infectionsincriticallyillsurgicalpatients.Ann.Surg.1994,220,751–758.
21. Pelz,R.K.;Lipsett,P.A.;Swoboda,S.M.;Diener‐West,M.;Hammond,J.M.;Hendrix,C.W.Thediagnostic
valueoffungalsurveillanceculturesincriticallyillpatients.Surg.Infect.(Larchmt)2000,1,273–281.
22. Arendrup,M.C.Epidemiologyofinvasivecandidiasis.Curr.Opin.Crit.Care2010,16,445–452.
23. Abi‐Said,D.;Anaissie,E.;Uzun,O.;Raad,I.;Pinzcowski,H.;Vartivarian,S.TheEpidemiologyof
HematogenousCandidiasisCausedbyDifferentCandidaSpecies.Clin.Infect.Dis.1997,24,1122–1128.
24. Trick,W.E.;Fridkin,S.K.;Edwards,J.R.;Hajjeh,R.A.;Gaynes,R.P.;NationalNosocomialInfections
SurveillanceSystemHospitalsSeculartrendofhospital‐acquiredcandidemiaamongintensivecareunit
patientsintheUnitedStatesduring1989‐1999.Clin.Infect.Dis.2002,35,627–630.
25. Guinea,J.;Zaragoza,Ó.;Escribano,P.;Martín‐Mazuelos,E.;Pemán,J.;Sánchez‐Reus,F.;Cuenca‐Estrella,
M.;CANDIPOPProject,GEIH‐GEMICOMED(SEIMC),andREIPIMolecularidentificationandantifungal
susceptibilityofyeastisolatescausingfungemiacollectedinapopulation‐basedstudyinSpainin2010and
2011.Antimicrob.AgentsChemother.2014,58,1529–1537.
26. Chow,J.K.;Golan,Y.;Ruthazer,R.;Karchmer,A.W.;Carmeli,Y.;Lichtenberg,D.;Chawla,V.;Young,J.;
Hadley,S.Factorsassociatedwithcandidemiacausedbynon‐albicansCandidaspeciesversusCandida
albicansintheintensivecareunit.Clin.Infect.Dis.2008,46,1206–1213.
27. Pfaller,M.A.;Diekema,D.J.;Turnidge,J.D.;Castanheira,M.;Jones,R.N.TwentyYearsoftheSENTRY
AntifungalSurveillanceProgram:ResultsforCandidaSpeciesFrom1997–2016.OpenForumInfect.Dis.
2019,6,S79–S94.
28. Pfaller,M.A.;Diekema,D.J.;InternationalFungalSurveillanceParticipantGroupTwelveyearsof
fluconazoleinclinicalpractice:globaltrendsinspeciesdistributionandfluconazolesusceptibilityof
bloodstreamisolatesofCandida.Clin.Microbiol.Infect.2004,10Suppl1,11–23.
29. Chassot,F.;Venturini,T.P.;Piasentin,F.B.;Rossato,L.;Fiorini,A.;Svidzinski,T.I.E.;Alves,S.H.Exploring
theInVitroResistanceofCandidaparapsilosistoEchinocandins.Mycopathologia2016,181,663–670.
30. Dudiuk,C.;Macedo,D.;Leonardelli,F.;Theill,L.;Cabeza,M.S.;Gamarra,S.;Garcia‐Effron,G.Molecular
ConfirmationoftheRelationshipbetweenCandidaguilliermondiiFks1pNaturallyOccurringAminoAcid
SubstitutionsandItsIntrinsicReducedEchinocandinSusceptibility.Antimicrob.AgentsChemother.2017,61,
e02644–e16.
31. Rex,J.H.;Pfaller,M.A.;Galgiani,J.N.;Bartlett,M.S.;Espinel‐Ingroff,A.;Ghannoum,M.A.;Lancaster,M.;
Odds,F.C.;Rinaldi,M.G.;Walsh,T.J.;etal.Developmentofinterpretivebreakpointsforantifungal
susceptibilitytesting:conceptualframeworkandanalysisofinvitro‐invivocorrelationdatafor
fluconazole,itraconazole,andcandidainfections.SubcommitteeonAntifungalSusceptibilityTestingof
theNationalCommitteeforClinicalLaboratoryStandards.Clin.Infect.Dis.1997,24,235–247.
32. Satoh,K.;Makimura,K.;Hasumi,Y.;Nishiyama,Y.;Uchida,K.;Yamaguchi,H.Candidaaurissp.nov.,a
novelascomycetousyeastisolatedfromtheexternalearcanalofaninpatientinaJapanesehospital.
Microbiol.Immunol.2009,53,41–44.
33. Eyre,D.W.;Sheppard,A.E.;Madder,H.;Moir,I.;Moroney,R.;Quan,T.P.;Griffiths,D.;George,S.;Butcher,
L.;Morgan,M.;etal.ACandidaaurisOutbreakandItsControlinanIntensiveCareSetting.N.Engl.J.
Med.2018,379,1322–1331.
J.Fungi2020,6,10112of14
34. Adams,E.;Quinn,M.;Tsay,S.;Poirot,E.;Chaturvedi,S.;Southwick,K.;Greenko,J.;Fernandez,R.;Kallen,
A.;Vallabhaneni,S.;etal.CandidaaurisinHealthcareFacilities,NewYork,USA,2013–2017. Emerg.Infect.
Dis.2018,24,1816–1824.
35. RuizGaitán,A.C.;Moret,A.;LópezHontangas,J.L.;Molina,J.M.;AleixandreLópez,A.I.;Cabezas,A.H.;
MollarMaseres,J.;Arcas,R.C.;GómezRuiz,M.D.;Chiveli,M.Á.;etal.NosocomialfungemiabyCandida
auris:FirstfourreportedcasesincontinentalEurope.Rev.Iberoam.Micol.2017,34,23–27.
36. Grim,S.A.;Berger,K.;Teng,C.;Gupta,S.;Layden,J.E.;Janda,W.M.;Clark,N.M.Timingofsusceptibility‐
basedantifungaldrugadministrationinpatientswithCandidabloodstreaminfection:correlationwith
outcomes.J.Antimicrob.Chemother.2012,67,707–714.
37. Fortún,J.;Martín‐Dávila,P.;Gómez‐GarcíadelaPedrosa,E.;Pintado,V.;Cobo,J.;Fresco,G.;Meije,Y.;
Ros,L.;Alvarez,M.E.;Luengo,J.;etal.Emergingtrendsincandidemia:Ahigherincidencebutasimilar
outcome.J.Infect.2012,65,64–70.
38. Clancy,C.J.;Nguyen,M.H.Findingthe“missing50%”ofinvasivecandidiasis:hownonculturediagnostics
willimproveunderstandingofdiseasespectrumandtransformpatientcare.Clin.Infect.Dis.2013,56,1284–
1292.
39. Pfeiffer,C.D.;Samsa,G.P.;Schell,W.A.;Reller,L.B.;Perfect,J.R.;Alexander,B.D.QuantitationofCandida
CFUinInitialPositiveBloodCultures.J.Clin.Microbiol.2011,49,2879–2883.
40. Willinger,B.;Kienzl,D.;Kurzai,O.DiagnosticsofFungalInfections.InHumanFungalPathogens;Kurzai,
O.,Ed.;TheMycota;Springer:Berlin/Heidelberg,Germany,2014;ISBN978‐3‐642‐39431‐7.
41. Bal,A.M.;McGill,M.RapidspeciesidentificationofCandidadirectlyfrombloodculturebrothsby
Sepsityper‐MALDI‐TOFmassspectrometry:Impactonantifungaltherapy.J.R.Coll.Phys.Edinb.2018,48,
114–119.
42. Salimnia,H.;Fairfax,M.R.;Lephart,P.R.;Schreckenberger,P.;DesJarlais,S.M.;Johnson,J.K.;Robinson,G.;
Carroll,K.C.;Greer,A.;Morgan,M.;etal.EvaluationoftheFilmArrayBloodCultureIdentificationPanel:
ResultsofaMulticenterControlledTrial.J.Clin.Microbiol.2016,54,687–698.
43. Simor,A.E.;Porter,V.;Mubareka,S.;Chouinard,M.;Katz,K.;Vermeiren,C.;Fattouh,R.;Matukas,L.M.;
Tadros,M.;Mazzulli,T.;etal.RapidIdentificationofCandidaSpeciesfromPositiveBloodCulturesbyUse
oftheFilmArrayBloodCultureIdentificationPanel.J.Clin.Microbiol.2018,56,doi:10.1128/JCM.01387–18.
44. Burnham,J.P.;Wallace,M.A.;Fuller,B.M.;Shupe,A.;Burnham,C.‐A.D.;Kollef,M.H.ClinicalEffectof
ExpeditedPathogenIdentificationandSusceptibilityTestingforGram‐NegativeBacteremiaand
CandidemiabyUseoftheAcceleratePhenoTMSystem.J.Appl.Lab.Med.2019,3,569–579.
45. Charnot‐Katsikas,A.;Tesic,V.;Love,N.;Hill,B.;Bethel,C.;Boonlayangoor,S.;Beavis,K.G.Useofthe
AcceleratePhenoSystemforIdentificationandAntimicrobialSusceptibilityTestingofPathogensin
PositiveBloodCulturesandImpactonTimetoResultsandWorkflow.J.Clin.Microbiol.2018,56,e01166‐
17.
46. Galiana,A.;Coy,J.;Gimeno,A.;Guzman,N.M.;Rosales,F.;Merino,E.;Royo,G.;Rodríguez,J.C.
EvaluationoftheSepsisFlowChipassayforthediagnosisofbloodinfections.PLoSONE2017,12,e0177627.
47. Hall,L.;Febre,K.M.L.;Deml,S.M.;Wohlfiel,S.L.;Wengenack,N.L.EvaluationoftheYeastTrafficLight
PNAFISHProbesforIdentificationofCandidaSpeciesfromPositiveBloodCultures.J.Clin.Microbiol.
2012,50,1446–1448.
48. Huang,T.‐D.;Melnik,E.;Bogaerts,P.;Evrard,S.;Glupczynski,Y.EvaluationoftheePlexBloodCulture
IdentificationPanelsforDetectionofPathogensinBloodstreamInfections.J.Clin.Microbiol.2019,57,
doi:10.1128/JCM.01597‐18.
49. Fuchs,S.;Lass‐Flörl,C.;Posch,W.DiagnosticPerformanceofaNovelMultiplexPCRAssayfor
CandidemiaamongICUPatients.J.Fungi.(Basel)2019,5,86.
50. Denina,M.;Scolfaro,C.;Colombo,S.;Calitri,C.;Garazzino,S.;BarbuiAnna,A.;Brossa,S.;Regina
MargheritaChildren’sHospitalBloodstreamInfectionsStudyGroupparticipants;Tovo,P.‐A.
Magicplex(TM)SepsisReal‐Timetesttoimprovebloodstreaminfectiondiagnosticsinchildren.Eur.J.
Pediatr.2016,175,1107–1111.
51. Zboromyrska,Y.;Cillóniz,C.;Cobos‐Trigueros,N.;Almela,M.;Hurtado,J.C.;Vergara,A.;Mata,C.;
Soriano,A.;Mensa,J.;Marco,F.;etal.EvaluationoftheMagicplexTMSepsisReal‐TimeTestfortheRapid
DiagnosisofBloodstreamInfectionsinAdults.Front.CellInfect.Microbiol.2019,9,56.
J.Fungi2020,6,10113of14
52. Schabereiter‐Gurtner,C.;Selitsch,B.;Rotter,M.L.;Hirschl,A.M.;Willinger,B.Developmentofnovelreal‐
timePCRassaysfordetectionanddifferentiationofelevenmedicallyimportantAspergillusandCandida
speciesinclinicalspecimens.J.Clin.Microbiol.2007,45,906–914.
53. Camp,I.;Manhart,G.;Schabereiter‐Gurtner,C.;Spettel,K.;Selitsch,B.;Willinger,B.Clinicalevaluationof
anin‐housepanfungalreal‐timePCRassayforthedetectionoffungalpathogens.Infection2020,48,345–
355.
54. Zeller,I.;Schabereiter‐Gurtner,C.;Mihalits,V.;Selitsch,B.;Barousch,W.;Hirschl,A.M.;Makristathis,A.;
Willinger,B.Detectionoffungalpathogensbyanewbroadrangereal‐timePCRassaytargetingthefungal
ITS2region.J.Med.Microbiol.2017,66,1383–1392.
55. Leitner,E.;Kessler,H.H.;Spindelboeck,W.;Hoenigl,M.;Putz‐Bankuti,C.;Stadlbauer‐Köllner,V.;Krause,
R.;Grisold,A.J.;Feierl,G.;Stauber,R.E.Comparisonoftwomolecularassayswithconventionalblood
culturefordiagnosisofsepsis.J.Microbiol.Methods2013,92,253–255.
56. Schreiber,J.;Nierhaus,A.;Braune,S.A.;deHeer,G.;Kluge,S.Comparisonofthreedifferentcommercial
PCRassaysforthedetectionofpathogensincriticallyillsepsispatients.Med.Klin.IntensivmedNotfmed
2013,108,311–318.
57. Wellinghausen,N.;Kochem,A.‐J.;Disqué,C.;Mühl,H.;Gebert,S.;Winter,J.;Matten,J.;Sakka,S.G.
DiagnosisofBacteremiainWhole‐BloodSamplesbyUseofaCommercialUniversal16SrRNAGene‐Based
PCRandSequenceAnalysis.J.Clin.Microbiol.2009,47,2759–2765.
58. Nieman,A.E.;Savelkoul,P.H.M.;Beishuizen,A.;Henrich,B.;Lamik,B.;MacKenzie,C.R.;Kindgen‐Milles,
D.;Helmers,A.;Diaz,C.;Sakka,S.G.;etal.Aprospectivemulticenterevaluationofdirectmolecular
detectionofbloodstreaminfectionfromaclinicalperspective.BMCInfect.Dis.2016,16,314.
59. Loonen,A.J.M.;deJager,C.P.C.;Tosserams,J.;Kusters,R.;Hilbink,M.;Wever,P.C.;vandenBrule,A.J.C.
Biomarkersandmolecularanalysistoimprovebloodstreaminfectiondiagnosticsinanemergencycare
unit.PLoSONE2014,9,e87315.
60. Neely,L.A.;Audeh,M.;Phung,N.A.;Min,M.;Suchocki,A.;Plourde,D.;Blanco,M.;Demas,V.;Skewis,
L.R.;Anagnostou,T.;etal.T2MagneticResonanceEnablesNanoparticle‐MediatedRapidDetectionof
CandidemiainWholeBlood.Sci.Transl.Med.2013,5,182ra54.
61. Mylonakis,E.;Clancy,C.J.;Ostrosky‐Zeichner,L.;Garey,K.W.;Alangaden,G.J.;Vazquez,J.A.;Groeger,
J.S.;Judson,M.A.;Vinagre,Y.‐M.;Heard,S.O.;etal.T2magneticresonanceassayfortherapiddiagnosis
ofcandidemiainwholeblood:aclinicaltrial.Clin.Infect.Dis.2015,60,892–899.
62. Arendrup,M.C.;Andersen,J.S.;Holten,M.K.;Krarup,K.B.;Reiter,N.;Schierbeck,J.;Helleberg,M.
DiagnosticPerformanceofT2CandidaAmongICUPatientsWithRiskFactorsforInvasiveCandidiasis.
Open.Forum.Infect.Dis.2019,6,ofz163.
63. Mylonakis,E.;Zacharioudakis,I.M.;Clancy,C.J.;Nguyen,M.H.;Pappas,P.G.EfficacyofT2Magnetic
ResonanceAssayinMonitoringCandidemiaafterInitiationofAntifungalTherapy:theSerialTherapeutic
andAntifungalMonitoringProtocol(STAMP)Trial.J.Clin.Microbiol.2018,56,e01756‐17.
64. Clancy,C.J.;Pappas,P.G.;Vazquez,J.;Judson,M.A.;Kontoyiannis,D.P.;Thompson,G.R.;Garey,K.W.;
Reboli,A.;Greenberg,R.N.;Apewokin,S.;etal.DetectingInfectionsRapidlyandEasilyforCandidemia
Trial,Part2(DIRECT2):AProspective,MulticenterStudyoftheT2CandidaPanel.Clin.Infect.Dis.2018,
66,1678–1686.
65. Zurl,C.;Prattes,J.;Zollner‐Schwetz,I.;Valentin,T.;Rabensteiner,J.;Wunsch,S.;Hoenigl,M.;Krause,R.
T2Candidamagneticresonanceinpatientswithinvasivecandidiasis:Strengthsandlimitations.Med.
Mycol.2019,2020,58,632‐638.
66. Sexton,D.J.;Bentz,M.L.;Welsh,R.M.;Litvintseva,A.P.EvaluationofanewT2MagneticResonanceassay
forrapiddetectionofemergentfungalpathogenCandidaaurisonclinicalskinswabsamples.Mycoses2018,
61,786–790.
67. Muñoz,P.;Vena,A.;Machado,M.;Gioia,F.;Martínez‐Jiménez,M.C.;Gómez,E.;Origüen,J.;Orellana,
M.Á.;López‐Medrano,F.;Fernández‐Ruiz,M.;etal.T2CandidaMRasapredictorofoutcomeinpatients
withsuspectedinvasivecandidiasisstartingempiricalantifungaltreatment:Aprospectivepilotstudy.J.
Antimicrob.Chemother.2018,73,iv6–iv12.
68. Muñoz,P.;Vena,A.;Machado,M.;Martínez‐Jiménez,M.C.;Gioia,F.;Gómez,E.;Origüen,J.;Orellana,
M.Á.;López‐Medrano,F.;Pérez‐Granda,M.‐J.;etal.T2MRcontributestotheveryearlydiagnosisof
complicatedcandidaemia.Aprospectivestudy.J.Antimicrob.Chemother.2018,73,iv13–iv19.
J.Fungi2020,6,10114of14
69. Clancy,C.J.;Nguyen,M.H.T2magneticresonanceforthediagnosisofbloodstreaminfections:Chartinga
pathforward.J.Antimicrob.Chemother.2018,73,iv2–iv5.
70. White,P.L.Recentadvancesandnovelapproachesinlaboratory‐baseddiagnosticmycology.Med.Mycol.
2019,57,S259–S266.
71. Pappas,P.G.;Kauffman,C.A.;Andes,D.R.;Clancy,C.J.;Marr,K.A.;Ostrosky‐Zeichner,L.ClinicalPractice
GuidelinefortheManagementofCandidiasis:2016UpdatebytheInfectiousDiseasesSocietyofAmerica.
Clin.InfectDis.Off.Publ.Infect.Dis.Soc.Am.2016,62,e1–e50.
72. Clancy,C.J.;Nguyen,M.H.Non‐CultureDiagnosticsforInvasiveCandidiasis:PromiseandUnintended
Consequences.J.Fungi.(Basel)2018,4,27.
73. Kidd,S.E.;Chen,S.C.‐A.;Meyer,W.;Halliday,C.L.ANewAgeinMolecularDiagnosticsforInvasive
FungalDisease:AreWeReady?Front.Microbiol.2020,10,2903.
©2020bytheauthors.LicenseeMDPI,Basel,Switzerland.Thisarticleisanopenaccess
articledistributedunderthetermsandconditionsoftheCreativeCommonsAttribution
(CCBY)license(http://creativecommons.org/licenses/by/4.0/).