Access to this full-text is provided by Wiley.
Content available from Oxidative Medicine and Cellular Longevity
This content is subject to copyright. Terms and conditions apply.
Research Article
Antiaging of Cucurbitane Glycosides from Fruits of
Momordica charantia L.
Xueli Cao, Yujuan Sun, Yanfei Lin, Yanjun Pan, Umer Farooq, Lan Xiang ,
and Jianhua Qi
College of Pharmaceutical Sciences, Zhejiang University, Yu Hang Tang Road 866, Hangzhou 310058, China
Correspondence should be addressed to Lan Xiang; lxiang@zju.edu.cn and Jianhua Qi; qijianhua@zju.edu.cn
Received 10 September 2017; Revised 21 December 2017; Accepted 11 January 2018; Published 25 March 2018
Academic Editor: Carolina G. Llorente
Copyright © 2018 Xueli Cao et al. This is an open access article distributed under the Creative Commons Attribution License,
which permits unrestricted use, distribution, and reproduction in any medium, provided the original work is properly cited.
Methanol extracts of Momordica charantia L. fruits are extensively studied for their antiaging activities. A new cucurbitane-type
triterpenoid (1) and nine other known compounds (2–10) were isolated, and their structures were determined according to their
spectroscopic characteristics and chemical derivatization. Biological evaluation was performed on a K6001 yeast bioassay
system. The results indicated that all the compounds extended the replicative lifespan of K6001 yeast significantly. Compound 9
was used to investigate the mechanism involved in the increasing of the lifespan. The results indicated that this compound
significantly increases the survival rate of yeast under oxidative stress and decreases ROS level. Further study on gene expression
analysis showed that compound 9 could reduce the levels of UTH1 and SKN7 and increase SOD1 and SOD2 gene expression. In
addition, it could not extend the lifespan of the yeast mutants of Uth1,Skn7,Sod1, and Sod2. These results demonstrate that
compound 9 exerts antiaging effects via antioxidative stress and regulation of UTH1,SKN7,SOD1, and SOD2 yeast gene expression.
1. Introduction
Fruits of Momordica charantia L. are edible healthy vegetable
in Asia and commonly known as bitter melon or bitter gourd
because of their bitter taste. Given their nutritional potential,
they are used as traditional Chinese herbal medicine to treat
several ailments, such as diabetes, constipation, abdominal
pain, kidney stones, piles, pneumonia, and improve appetite
[1–5]. M. charantia contains biologically active phytochemi-
cals, such as polysaccharides, proteins, flavonoids, glycosides,
saponins, steroids, alkaloids, essential oils, and triterpenes
[5–10]. Many of these phytochemicals exhibit antitumor,
anti-inflammatory, immunomodulation, and antidiabetic
activities and the ability to reduce oxidative stress [5].
Aging is a dominating risk factor for age-related diseases,
including cancer, metabolic disease, cardiovascular disease,
and neurodegenerative illnesses [11]. As the aging popula-
tion is increasing dramatically throughout the world, aging
has drawn great attention because of huge expenses for
medical care and serious consequences of the related dis-
eases. Interventions that delay aging were found to have
a greater effect on the quality of life compared with
disease-specific approaches [12]. In our previous studies
[13–17], a yeast mutant K6001 was employed in the bioassay
system, and ganodermasides A–D, phloridzin, nolinospiro-
side F, and parishin with significant antiaging potential from
natural sources were obtained.
Basing on the K6001 bioassay system, we isolated
one novel cucurbitane glycoside (1) and nine known
cucurbitane-type triterpenoids (2–10) from the fruits of
M. charantia L. (Figure 1). Essential studies on the action
mechanism suggested that these cucurbitane glycosides could
improve the antioxidative properties of yeasts. The yeast
genes of youth 1 (UTH1), skinhead-7 (SKN7), and superox-
ide dismutase (SOD) may also be involved in the action.
2. Material and Methods
2.1. General. The chemical reagents used were of HPLC
grade and purchased from TEDIA (Rhode Island, USA).
The others were of analytical grade and obtained from
Sinopharm Chemical Reagent Co. Ltd. (Shanghai, China).
Hindawi
Oxidative Medicine and Cellular Longevity
Volume 2018, Article ID 1538632, 10 pages
https://doi.org/10.1155/2018/1538632
The preparative HPLC system was equipped with two
ELITE P-230 pumps and an UV detector. Optical rotations
were determined on a JASCO P-1030 digital polarimeter.
High-resolution ESI-TOF-MS analyses were performed on
an Agilent Technologies 6224A Accurate-Mass TOF LC/
MS system (Santa Clara, CA, USA). Nuclear magnetic reso-
nance (NMR) spectra were recorded on a Bruker AV III-500
spectrometer (Bruker, Billerica, MA, USA). Column chroma-
tography was performed over the silica gel (200–300 mesh,
Yantai Chemical Industry Research Institute, Yantai, China)
or reversed phase C18 (Octadecylsilyl, ODS) silica gel
(Cosmosil 75C
18
-OPN, Nacalai Tesque, Japan).
2.2. Plant Material and Yeast Strains. Fruits of M. charantia
were purchased from Liangzhu market of Hangzhou,
Zhejiang Province, China, 2011. The identity of this plant
was confirmed by an associate professor Liurong Chen, and
a voucher specimen (number 20110712) was preserved in
Zhejiang University Institute of Materia Medica. The yeast
strains BY4741 and mutants of uth1,skn7,sod1, and sod2
with K6001 background are from Prof. Matsuura in Chiba
University, and K6001 strain with W303 background is from
Prof. Breitenbach in Salzburg University.
2.3. Extraction and Isolation. About 1.5 kg (dry weight) of the
material was smashed and extracted with methanol (MeOH)
for 3 days with shaking at room temperature. The extract was
filtered and concentrated to obtain a crude extract (224 g),
which was partitioned with ethyl acetate (EtOAc) and water.
The active EtOAc layer (30 g) was subjected to a silica gel
open column with n-hexane/acetone (99 : 1, 98 : 2, 95 : 5,
90 : 10, 80 : 20, 70 : 30, 60 : 40, 50 : 50, 20 : 80, and 0 : 100) and
acetone/MeOH (50 : 50 and 0 : 100). The EtOAc layer yielded
nine fractions after the combination based on TLC analysis.
The eighth active fraction of 4.0 g (13.9g in total) was subse-
quently separated by ODS open column with MeOH/H
2
O
(60 : 40, 70 : 30, 75 : 25, 80 : 20, 90 : 10, and 100 : 0), and 11
samples were obtained (fr.1–fr.11). The four active samples
of fr.6–fr.9 were further separated as follows:
Fr.6 (600 mg) was introduced to an ODS open column
with MeOH/H
2
O (70 : 30, 73 : 27, 75 : 25, 77 : 23, 80 : 20,
83 : 17, 85 : 15, and 100 : 0) to yield nine samples (fr.6-1–
fr.6-9). Fr.6-6 (72 mg) was further separated by a silica
gel open column chromatography with dichloromethane
(CH
2
Cl
2
)/MeOH (100 : 0, 99 : 1, 98 : 2, 97.5 : 2.5, 95 : 5, 90 : 10,
and 0 : 100), and the sixth active fraction (11.6 mg) was
purified by HPLC [C30-UG-5 (φ10 ×250 mm, Nomura
Chemical), mobile phase: acetonitrile (MeCN)/H
2
O (90 : 10),
flow rate: 3 mL/min, and detector: 210 nm] and yielded
compound 1 (4.6 mg, t
R
= 33.5 min) and compound 2
(1.0 mg, t
R
= 30.7 min).
Fr.7 (277 mg) was separated in a silica gel open column
with chloroform (CHCl
3
)/MeOH (100 : 0, 98 : 2, 95 : 5,
90 : 10, and 0 : 100) and yielded fr.7-1–fr.7-11. Subsequently,
fr.7-7 (63 mg) was subjected to ODS open column chroma-
tography with MeOH/H
2
O (70 : 30, 75 : 25, 80 : 20, 90 : 10,
and 100 : 0), and the second fraction (37.8 mg) was purified
by HPLC [C30-UG-5 (φ10 ×250 mm, Nomura Chemical),
mobile phase: MeCN/H
2
O (62 : 38), flow rate: 3 mL/min,
and detector: 210 nm] and yielded compound 3 (10.6 mg,
t
R
= 15.9 min), comp ound 4 (1.8 mg, t
R
= 17.1 min), and
compound 5 (1.9 mg, t
R
= 18.7 min).
Fr.8 (340 mg) was introduced to a silica open column
with n-hexane/CHCl
3
(50 : 50, 30 : 70, and 0 : 100) and
CHCl
3
/MeOH (97 : 3, 95 : 5, 90 : 10, and 0 : 100) to give
fr.8-1–fr.8-9. Then, fr.8-4 (49 mg) was purified by HPLC
18
12 R
17
14
O
19
2
AllO 4
O
730
AllO AllO
5
O
OH
O
R
21
R= 23
26
OH
1R=
OH
2
3
6
9
O
O
4
7
O
O
O
8
10
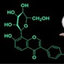
Figure 1: Chemical structures of compounds 1–10.
2 Oxidative Medicine and Cellular Longevity
[CAPCELL PAKC
18
, Shiseido (φ10 ×250 mm), mobile phase:
MeCN/H
2
O (63 : 37), flow rate: 3 mL/min, and detector:
210 nm] and yielded compound 6 (10.2 mg, t
R
= 20.2 min),
compound 7 (4.9 mg, t
R
= 22.9 min), and compound 8
(1.3 mg, t
R
= 32.0 min).
Fr.9 (600 mg) was separated by silica gel open column
with CHCl
3
/MeOH (100 : 0, 100 : 1, 100 : 2, 100 : 3, 100 : 5,
90 : 10, and 0 : 100), and fr.9-1–fr.9-8 were obtained. Then,
fr.9-4 (332 mg) was further separated by ODS open column
with MeOH/H
2
O (90 : 10, 95 : 5, and 100 : 0), and the
third fraction (42 mg) was purified by HPLC [CAPCELL
PAKC
18
, Shiseido (φ10 ×250 mm), mobile phase: MeCN/
H
2
O (60 : 40), flow rate: 3 mL/min, and detector: 210 nm]
and yielded compound 9 (5.0 mg, t
R
= 28.7 min) and com-
pound 10 (8.0 mg, t
R
= 33.8 min).
2.3.1. Compound 1. White solid; α20
D-78.6 (c0.2, CH
3
OH);
high-resolution ESI-TOF-MS m/z653.4029, calculated
for C
37
H
58
O
8
Na (M + Na)
+
653.4024; data of
1
H NMR
and
13
C NMR are shown in Table 1.
2.3.2. Charantoside IV (2). Colorless solid; α28
D-143.9
(c0.16, CH
3
OH); MS m/z623 (M + Na)
+
,C
36
H
56
O
7
Na;
13
C NMR (125 MHz, pyridine-d
5
): δ= 15.4, 19.3, 19.3,
19.3, 20.6, 21.5, 24.2, 26.0, 28.0, 28.7, 31.4, 33.8, 37.2,
39.4, 40.5, 40.5, 45.7, 45.9, 49.3, 51.0, 52.7, 63.7, 69.7,
72.9, 73.5, 76.6, 80.5, 85.5, 86.3, 104.3, 115.1, 130.3,
130.4, 134.6, 135.1, and 142.9. The structure was identified
based on comparison of MS,
1
H, and
13
C NMR data with
literature [18].
2.3.3. Momordicoside F
2
(3). White solid; α20
D-101.0
(c0.94, CHCl
3
:CH
3
OH = 1 : 1); MS m/z641 (M + Na)
+
,
C
36
H
58
O
8
Na;
13
C NMR (125 MHz, pyridine-d
5
): δ= 15.4,
19.3, 19.3, 20.6, 21.5, 24.3, 26.0, 28.0, 28.6, 31.3, 31.3,
31.4, 33.8, 37.0, 39.4, 39.9, 40.5, 45.7, 45.8, 49.3, 50.6,
52.7, 63.7, 69.7, 70.1, 72.9, 73.5, 76.6, 80.5, 85.5, 86.3,
104.2, 124.6, 130.4, 134.6, and 142.1. The structure was
identified based on comparison of MS,
1
H, and
13
C NMR
data with literature [19].
2.3.4. Goyaglycoside-b (4). White solid; α20
D-100.7 (c0.2,
CH
3
OH); MS m/z671 (M + Na)
+
,C
37
H
60
O
9
Na;
13
C NMR
(125 MHz, pyridine-d
5
): δ= 15.3, 19.1, 19.3, 20.4, 21.7, 23.7,
25.3, 27.8, 28.6, 31.3, 31.3, 31.3, 34.3, 37.0, 39.5, 40.0, 42.0,
42.6, 45.7, 48.6, 48.7, 50.8, 58.1, 63.7, 69.7, 70.1, 72.2, 74.2,
77.0, 83.9, 86.0, 102.8, 112.8, 124.7, 132.0, 133.6, and 142.1.
The structure was identified based on comparison of MS,
1
H, and
13
C NMR data with literature [4].
2.3.5. Karaviloside III (5). White solid; α28
D+ 70.9 (c0.1,
CH
3
OH); MS m/z657 (M + Na)
+
,C
37
H
62
O
8
Na;
13
C NMR
(125 MHz, pyridine-d
5
): δ= 16.0, 18.5, 19.4, 23.1, 26.3, 28.3,
29.3, 29.4, 29.6, 30.8, 31.3, 31.3, 33.2, 34.8, 35.4, 37.1, 39.8,
40.0, 42.4, 46.7, 48.7, 49.3, 50.6, 56.6, 63.8, 69.7, 70.1, 72.5,
73.9, 76.1, 78.0, 88.3, 105.4, 119.5, 124.2, 142.2, and 148.4.
The structure was identified based on comparison of MS,
1
H, and
13
C NMR data with literature [20].
Table 1:
1
H-NMR (500 MHz) and
13
C-NMR (125 MHz) data of
compound 1 in pyridine-d
5
.
Position Compound 1
δ
H
(Jin Hz) δ
C
1α1.46 19.2
1β1.91 —
2α1.75 27.8
2β2.17 —
3α3.73 (br s) 83.9
4—39.5
5—86.0
6 6.18 (dd, 2.0, 9.7) 133.6
7 5.63 (dd, 3.6, 9.7) 132.0
8β3.15 (br s) 42.7
9—48.6
10α2.48 (dd, 5.6, 12.7) 42.1
11α1.75 23.7
11β1.68 —
12α1.60 31.3
12β1.55 —
13 —45.8
14 —48.7
15α1.31 34.3
15β1.31 —
16α1.94 28.7
16β1.32 —
17α1.50 51.2
18 0.91 (s) 15.3
19 4.91 (s) 112.8
20 1.50 37.3
21 0.95 (d, 5.6) 19.4
22α1.83 40.5
22β2.32 —
23 5.76 (m) 130.4
24 6.32 (d, 15.3) 135.1
25 —143.0
26α4.96 (s) 115.1
26β5.10 (s) —
27 1.92 (s) 19.3
28 0.83 (s) 25.3
29 1.47 (s) 21.7
30 0.90 (s) 20.4
−OCH
3
3.52 (s) 58.1
1´ 5.52 (d, 7.8) 102.8
2´ 3.94 (dt, 2.6, 7.6) 74.2
3´ 4.76 (d, 2.8) 72.2
4´ 4.23 (td, 2.8, 9.3) 69.7
5´ 4.50 (m) 77.0
6´α4.43 (m) 63.7
6´β4.57 (m) —
3Oxidative Medicine and Cellular Longevity
2.3.6. Charantoside VI (6). White solid; α28
D-67.2 (c0.48,
CH
3
OH); MS m/z655 (M + Na)
+
,C
37
H
60
O
8
Na;
13
C NMR
(125 MHz, pyridine-d
5
): δ= 15.2, 18.8, 19.3, 20.3, 20.6, 21.5,
24.2, 26.0, 26.3, 28.0, 29.1, 31.5, 33.8, 34.2, 39.4, 40.5, 43.4,
45.7, 45.9, 49.3, 51.7, 52.7, 55.7, 63.7, 69.7, 72.9, 73.5, 76.6,
76.8, 80.5, 85.6, 86.3, 104.2, 127.8, 130.4, 134.5, and 135.5.
The structure was identified based on comparison of MS,
1
H, and
13
C NMR data with literature [18].
2.3.7. Charantagenin E (7). Colorless solid; α28
D-104.3
(c0.11, CH
3
OH); MS m/z685 (M + Na)
+
,C
38
H
62
O
9
Na;
13
C NMR (125 MHz, pyridine-d
5
): δ= 15.1, 18.8, 19.1,
20.4, 20.4, 21.7, 23.7, 25.3, 26.2, 27.8, 29.0, 31.4, 34.2,
34.2, 39.5, 42.0, 42.6, 43.4, 45.7, 48.5, 48.7, 51.8, 55.7,
58.0, 63.7, 69.7, 72.2, 74.2, 76.9, 77.0, 83.9, 86.0, 102.8,
112.8, 127.8, 132.0, 133.5, and 135.5. The structure was
identified according to the comparison among MS,
1
H,
and
13
C NMR data in literature [21].
2.3.8. Charantoside II (8). White solid; α20
D-67.1 (c0.2,
CH
3
OH); MS m/z685 (M + Na)
+
,C
38
H
62
O
9
Na;
13
C NMR
(125 MHz, pyridine-d
5
): δ= 15.2, 18.5, 19.1, 19.3, 20.4, 21.6,
23.7, 25.2, 26.2, 27.7, 28.8, 31.4, 33.2, 34.2, 39.5, 42.0, 42.6,
43.7, 45.8, 48.5, 48.7, 51.7, 56.0, 58.0, 63.6, 69.6, 72.2, 74.1,
75.2, 76.9, 84.0, 85.9, 102.9, 112.8, 128.3, 132.0, 133.5, and
134.9. The structure was identified based on comparison of
MS,
1
H, and
13
C NMR data with literature [18].
2.3.9. Momordicoside G (9). White solid; α28
D-90.2 (c1.0,
CH
3
OH); MS m/z655 (M + Na)
+
,C
37
H
60
O
8
Na;
13
C NMR
(125 MHz, pyridine-d
5
): δ= 15.5, 19.3, 20.6, 21.5, 24.3, 26.0,
26.5, 26.9, 28.0, 28.6, 31.5, 33.8, 36.7, 39.4, 40.1, 40.5, 45.7,
45.9, 49.3, 50.6, 50.7, 52.7, 63.7, 65.3, 69.7, 72.9, 73.5, 75.3,
76.6, 80.6, 85.5, 86.3, 104.2, 128.8, 130.4, 134.6, and 138.1.
The structure was identified based on comparison of MS,
1
H, and
13
C NMR data with literature [19].
2.3.10. Goyaglycoside-d (10). White solid; α28
D-124.9 (c0.1,
CH
3
OH); MS m/z685 (M + Na)
+
,C
38
H
62
O
9
Na;
13
C NMR
(125 MHz, pyridine-d
5
): δ= 15.3, 19.1, 19.3, 20.4, 21.6, 23.7,
25.3, 26.4, 26.9, 27.8, 28.6, 31.3, 34.3, 36.8, 39.5, 40.1, 42.0,
42.6, 45.7, 48.5, 48.7, 50.5, 50.8, 58.1, 63.7, 69.6, 72.2, 74.1,
75.3, 77.0, 83.9, 86.0, 102.8, 112.8, 128.9, 132.0, 133.6, and
138.0. The structure was identified based on comparison of
MS,
1
H, and
13
C NMR data with literature [4].
2.4. Acid Hydrolysis and Sugar Analysis of Compound 1. The
absolute configuration of sugar moiety of compound 1 was
determined according to a previously reported method
[22, 23]. Briefly, compound 1 (0.5 mg) in anhydrous 2.0 M
HCl in MeOH (1 mL) was heated at 80
°
C with reflux for
4 h. The reaction solution was evaporated and partitioned
between chloroform and water. The residue of aqueous layer
was heated with 0.5 mg L-cysteine methyl ester in pyridine
(200 μL) at 60
°
C for 1 h; then, o-tolyl isothiocyanate dissolved
in 100 μL pyridine (7 mg/mL) was added to the reaction
mixture and further reacted at 60
°
C for 1 h. After that, the
reaction mixture was dried and analyzed by LC/HRESIMS
with the following conditions: Agilent Extend C18 column
(3.5 μm, 3.0 ×100 mm); DAD detection, 210 nm; t=0min
CH
3
OH/H
2
O/formic acid (30 : 70 : 0.1), t=15min CH
3
OH/
H
2
O/formic acid (60 : 40 : 0.1); and flow rate: 0.45 mL/min.
The allose thiocarbamate standards were prepared in the
same procedure. Given that L-allose is limitedly available, the
retention time of L-allose thiocarbamate derivative was
obtained by reacting D-allose with D-cysteine methyl ester.
The basis of this approach is the fact that the t
R
values of
D- and L-enantiomers are reversed when D-cysteine methyl
ester is used [22].
2.5. Lifespan Assay. The bioassay method was performed as
described in a previous study [13]. Briefly, K6001 or mutants
with K6001 background were grown on a YPGalactose
medium consisting of 3% galactose, 2% hipolypeptone, and
1% yeast extract or on a YPGlucose medium containing 2%
glucose instead of galactose. Agar plates were prepared by
adding 2% agar to the medium. For screening, the K6001
yeast strain was first incubated in the galactose medium for
24 h with shaking and then centrifuged. The yeast pellet
was washed with PBS three times. The cells were then diluted
and counted using a hemocytometer, and approximately
4000 cells were plated on glucose agar plates containing
different concentrations of samples. The plates were stored
in an incubator at 28
°
C. After 48 h, the yeast cells in the plates
were observed with a microscope. For each plate, 40 colonies
were selected randomly, and the number of their daughter
cells was counted and analyzed.
2.6. Antioxidative Stress Method. Antioxidative stress assay
was performed as previously described with minor modi-
fication [16]. BY4741 yeast was inoculated in 5 mL of
YPGlucose medium and cultured at 28
°
C with shaking
for 24 h. The yeast cells at 0.1 OD
600
were transferred in
20 mL of new YPGlucose medium and incubated with com-
pound 9 at 1 and 3 μM or resveratrol (Res, positive control)
at 10 μM for 12 h.
For the first method, 5 μL aliquot after double dilution
from each group was dropped in the same YPGlucose agar
plate mixed with 9 mM H
2
O
2
, and the plate was incubated
at 28
°
C for 4 days. The growth rates of the yeast cells in
different groups were compared and photographed.
Another antioxidative stress assay was used to validate
the accuracy of the experiment. Approximately 200 cells
mixed with the test samples were spreaded on YPGlucose
agar plates with or without 5 mM H
2
O
2
and cultured at
28
°
C for 48 h. The survival rates of the sample groups were
counted and compared with those of the control group.
2.7. Determination of ROS Level in Yeast. The ROS assay
procedure was the same with a previous study [17]. BY4741
yeast cells were cultured as described in the experiment above
and incubated with compound 9 at 1 or 3 μM for 23 h.
Changes in intracellular ROS levels of the yeast were deter-
mined using an ROS assay kit (Beyotime, Jiangsu, China)
and a fluorescent plate reader (Spectra Max M2, Molecular
Devices, San Francisco, CA, USA). A total of 1 mL of cultured
broth was obtained, treated with 10 μM DCFH-DA at 28
°
Cin
dark, and then shaken by vortexing at 160 rpm at 15 min
intervals for 1 h. The yeast cells were subsequently washed
4 Oxidative Medicine and Cellular Longevity
with PBS, and their DCF fluorescence was measured by a
fluorescent plate reader at excitation and emission wave-
lengths of 488 and 525 nm, respectively.
2.8. Real-Time Quantitative PCR Analysis. BY4741 yeast
cells were cultured in glucose medium following the addi-
tion of 0, 1, 3 μM compound 9. RNA was extracted from
yeast cells in the exponential phase through the hot-
phenol method. Reverse transcription was performed using
a cloned AMV first-strand cDNA synthesis kit (Invitrogen,
California, USA) with oligo (dT) adaptor primers and 5 μg
of yeast total RNA. Real-time PCR was performed using
the CFX96-Touch (Bio-Rad, Hercules, USA) and SYBR
Premix EX Taq™(TaKaRa, Otsu, Japan). Thermal cycling
parameters for UTH1 and SKN7: 40 cycles, 94
°
C for 15 s,
55.4
°
C for 15 s, and 68
°
C for 20 s; for SOD1 and SOD2:
40 cycles, 94
°
C for 15 s, 60
°
C for 25 s, and 72
°
C for 10 s.
Primers used were as follows: for UTH1, sense 5′-CGC
CTC TTC CTC TT-3′and antisense 5′-ACC ATC GGA
AGG TTG TTC AG-3′; for SKN7, sense 5′-AGT TGT CAG
CGA CGG TCT TT-3′and antisense 5’-GCT GTG GCA
CCA TCT AGG TT-3′; for SOD1 sense 5′-CAC CAT TTT
CGT CCG TCT TT-3′and antisense 5′-TGG TTG TGT
CTC TGC TGG TC-3′; for SOD2, sense 5′-CTC CGG TCA
AAT CAA CGA AT-3′and antisense 5′-CCT TGG CCA
GAA GAT CTG AG-3′; for TUB1, sense 5′-CCA AGG
GCT ATT TAC GTG GA-3′and antisense 5′-GGT GTA
ATG GCC TCT TGC AT-3′. The amount of UTH1,SKN7,
SOD1, and SOD2 was normalized to that of TUB1.
2.9. Statistical Analysis. One-way analysis of variance was
performed using GraphPad Prism biostatistics software
(San Diego, CA, USA) to analyze the data. Significant
differences were compared by two-tailed multiple t-tests
with Student–Newman–Keuls test. Data were expressed
as means ±SEM of triplicate experiments. A P<0 05 was
considered statistically significant.
3. Results and Discussion
3.1. Structure Elucidation of Compound 1. Compound 1
has the molecular formula C
37
H
58
O
8
as determined by
HR-ESIMS measurement. The
1
H NMR data showed six
methyl groups at δ
H
0.83 (3H, s), 0.90 (3H, s), 0.91 (3H, s),
0.95 (3H, d, J=5 6Hz), 1.47 (3H, s), and 1.92 (3H, s), along
with six olefinic protons at δ
H
4.96 (1H, s), 5.10 (1H, s), 5.63
(1H, dd, J=3 6, 9.7 Hz), 5.76 (1H, m), 6.18 (1H, dd, J=210,
9.7 Hz), and 6.32 (1H, d, J=15 3Hz). Several multiple peaks
at δ
H
3.94–4.76 and the signal of an anomeric proton [δ
H
5.52 (1H, d, J=7 8Hz)] indicated the existence of a sugar
moiety. The
13
C NMR data revealed the presence of 37 car-
bon signals. With the combined signals of
13
C NMR and
DEPT, the 37 carbon signals were attributed to six olefinic
carbons (δ
C
115.1, 130.4, 132.0, 133.6, 135.1, and 143.0),
one anomeric carbon (δ
C
102.8), one oxygenated quaternary
carbon (δ
C
86.0), six oxymethines (δ
C
69.7, 72.2, 74.2, 77.0,
83.9, and 112.8), one oxymethylene (δ
C
63.7), one methoxy
group (δ
C
58.1), four quaternary sp
3
carbons (δ
C
39.5, 45.8,
48.6, and 48.7), four methines (δ
C
37.3, 42.1, 42.7, and
51.2), seven methylenes (δ
C
19.2, 23.7, 27.8, 28.7, 31.3, 34.3,
and 40.5), and six methyl groups (δ
C
15.3, 19.3, 19.4, 20.4,
21.7, and 25.3). Detailed analysis of the
1
H-
1
H COSY spectra
led to the determination of the partial structures depicted by
the bonds (Figure 2, in bold bonds). In the HMBC spectrum,
these partial structures were connected to yield the following
gross structures: H-3 to C-5; H-6 to C-5; H-8 to C-9; H-10 to
C-9; CH
3
-18 to C-12, C-13, C-14, and C-17; H-19 to C-9,
C-10, C-11, and −OCH
3
;CH
3
-28 to C-3, C-4, and C-29;
CH
3
-29 to C-4, C-5, and C-28; CH
3
-30 to C-8, C-13, C-14,
and C-15; H-23 to C-25; H-24 to C-25, C-26, and C-27;
H-26 to C-24; CH
3
-27 to C-24; and −OCH
3
to C-19. The
signals of H-3 to C-1′of allose and anomeric proton H-1′
of allose to C-3 in the HMBC indicated the location of
the sugar moiety (Figure 2). The βanomeric configuration
of allose was determined from its coupling constant J
(7.8 Hz) of anomeric protons (δ
H
5.52). The absolute con-
figuration of the sugar moiety was further confirmed by
the degradation of compound 1 and through the comparison
of the retention time of its aldose thiocarbamate derivative
(t
R
= 9.287 min) with those of the following aldose thiocar-
bamate standards: L-cysteine-D-allose (t
R
= 9.127 min) and
D-cysteine-D-allose (t
R
= 7.460 min).
The relative stereochemistry of compound 1 was deduced
by nuclear overhauser enhancement spectroscopy (NOESY)
analysis. As shown in Figure 2, the broad singlet signal
of H-3 appeared at δ
H
3.73 thereby suggested the αcon-
figuration of this proton. The NOESY correlations of
HO
HO
OO
2
1
11
21
20
26
15
30
6
28
COSY
HMBC
O
OH
HO
O
AllO
H
O
H
H
H
OMe
H
H
NOESY
H
H
Figure 2: Gross structure of compound 1 with
1
H-
1
H COSY, selected HMBC, and NOESY correlations.
5Oxidative Medicine and Cellular Longevity
0
3
6
9
12
1 M
e yeast generations of K6001
⁎⁎⁎
⁎⁎⁎⁎
⁎⁎⁎
⁎⁎
⁎⁎⁎
⁎⁎⁎ ⁎⁎⁎⁎⁎⁎
⁎⁎ ⁎⁎⁎ ⁎⁎ ⁎⁎ ⁎⁎ ⁎⁎ ⁎⁎ ⁎⁎⁎ ⁎⁎ ⁎⁎⁎
Control Res C1 C2 C3 C4 C5 C6 C7 C8 C9 C10
3 M
Figure 3: Effect of compounds 1–10 on the replicative lifespan of K6001 yeast strain. The average lifespan of K6001 was as follows:
control (7.70 ±0.48); Res at 10 μM (10.20 ±0.42∗∗∗); compound 1 at 1 μM (9.20 ±0.49∗) and at 3 μM (9.25 ±0.42∗); compound 2 at
1μM (9.03 ±0.43∗) and at 3 μM (9.00 ±0.42∗); compound 3 at 1 μM (11.03 ±0.53∗∗∗) and at 3 μM (9.68 ±0.55∗∗); compound 4 at
1μM (11.08 ±0.50∗∗∗ ) and at 3 μM (10.25 ±0.56∗∗∗ ); compound 5 at 1 μM (10.40 ±0.45∗∗∗ ) and at 3 μM (10.70 ±0.45∗∗∗ ); compound 6
at 1 μM (9.68 ±0.57∗∗) and at 3 μM (10.10 ±0.46∗∗∗ ); compound 7 at 1 μM (9.68 ±0.55∗∗ ) and at 3 μM (9.75 ±0.55∗∗); compound 8 at
1μM (9.63 ±0.52∗∗ ) and at 3 μM (9.73 ±0.55∗∗); compound 9 at 1 μM (9.75 ±0.47∗∗) and at 3 μM (10.13 ±0.41∗∗∗); and compound 10 at
1μM (9.55 ±0.42∗∗) and at 3 μM (10.08 ±0.39∗∗∗ )(
∗P<005,∗∗ P<0 01, and ∗∗∗ P<0 001, compared with the control).
Control
Res
(10 M)
9
(1 M)
9
(3 M)
(a)
0
20
40
60
80
100
Control 31
9 (M)
Survival rate under
oxidative condition (%)
⁎⁎⁎
Res (10 M)
(b)
0.0
0.5
1.0
1.5
2.0
2.5
3.0
3.5
Control Res (10 M) 1
9 (M)
Fluorescence/107 cells
⁎⁎ ⁎⁎ ⁎
3
(c)
Figure 4: Effect of compound 9 on the antioxidative ability of yeast cells and ROS level of yeast. (a) BY4741 yeast cells in the control group
and compound 9-treated groups were dropped in the same YPGlucose agar plate mixed with 9 mM H
2
O
2
. After four days, the growth of yeast
cells in different groups was photographed. (b) Effect of compound 9 on the survival rates of yeast under oxidative stress condition. Control,
48.31 ±4.94; Res, 72.74 ±6.19∗; compound 9 at 1 μM, 71.21 ±2.75∗and at 3 μM, 68.73 ±5.89∗. The experiment was conducted at least thrice.
Vertical bars represent the mean ±SEM of three assays (∗P<0 05). (c) The change of ROS level of yeast after administration compound 9 at 1,
3μM. Control, 2.77 ±0.15∗∗; Res, 1.79 ±0.15∗∗; compound 9 at 1 μM, 1.98 ±0.08∗∗, and at 3 μM, 1.73 ±0.22∗. Vertical bars represent the
mean ±SEM of 6 repeats (∗P<005 and ∗∗ P<0 01, compared with the control).
6 Oxidative Medicine and Cellular Longevity
major cross-peaks of H-3α/Me-28, Me-28/H-10, H-10/Me-
30, and Me-30/H-17 indicated the α-orientation of these
protons. The correlations between H-1β/H-19, H-19/H-8,
H-8/Me-18, and Me-18/H-20 suggested the β-orientation
of these groups. The double bond at C-23 and C-24 was elu-
cidated by COSY correlations, whereas the transgeometry
was determined from the coupling constant, J
23-24
= 15.3 Hz.
The above evidence suggested that compound 1 was
structurally similar to (19R,23E)-5β, 19-epoxy-19-meth-
oxycucurbita-6, 23-25-trien-3β-ol 3-O-β-D-allopyranoside
(Figure 1).
3.2. Identification of the Known Compounds. Compounds
2–10 (Figure 1) were identified by comparing their spec-
troscopic data with those in literature.
3.3. Antiaging Activity in K6001 Yeast Strain. All isolated
cucurbitane triterpenoids (1–10)were tested for antiaging
activity through the K6001 bioassay method at different
optimum concentrat ions. All the compounds at 1 and 3 μM
extended the replicative lifespan of K6001 significantly
(Figure 3), demonstrating that the cucurbitane triterpenoids
isolated from M. charantia L. fruits have antiaging effect
in yeast.
3.4. Compound 9 Improves the Oxidative Resistance and
Decreases ROS Production of Yeast. Studies on mechanism
of action were conducted with compound 9 because of its
abundance and good activity. Oxidative stress is one of the
primary causes of aging, as indicated in various model
organisms [24]. Therefore, the effect of compound 9 on the
oxidative resistance of yeast was first tested. The growth of
yeast cells was inhibited at 9 mM H
2
O
2
, whereas incubation
with compound 9 at 1 or 3 μM remitted the inhibition
(Figure 4(a)). The effect was further confirmed in another
assay. As shown in Figure 4(b), the survival rate of the
control group was 48.31% ±4.94%, whereas that in the
experimental groups increased to 71.21% ±2.75% (com-
pound 9 at 1 μM, P<05) and 68.73% ±5.89% (compound
9at3μM, P<05). The experiments indicated that com-
pound 9 enhances the oxidative resistance of yeast cells.
Furthermore, we detected the ROS level of yeast after admin-
istration compound 9 at 1 and 3 μM. As we expected, the
ROS level of yeast in the resveratrol and compound 9 groups
were significantly decreased compared with the control
0.0
0.6
1.2
1.8
9 (M)
12 h
Relative level of SOD1 mRNA
Control Res (10 M) 13
⁎
⁎⁎
(a)
0.0
0.6
1.2
1.8
Control Res (10 M) 13
9 (M)
12 h
Relative level of SOD2 mRNA
⁎
⁎
(b)
0.0
0.6
1.2
1.8
9 (M)
24 h
Relative level of UTH1 mRNA
Control Res (10 M) 13
⁎⁎
⁎
⁎⁎⁎
(c)
0.0
0.6
1.2
1.8
9 (M)
24 h
Relative level of SKN7 mRNA
Control Res (10 M) 13
⁎
⁎
⁎
(d)
Figure 5: Effects of compound 9 on SOD1 (a), SOD2 (b), UTH1 (c), and SKN7 (d) yeast gene expression. The gene levels of BY4741 yeast cells
were tested after treated with compound 9 at 1 and 3 μM. Compound 9 significantly increased SOD1 and SOD2 yeast gene level at 12 h and
inhibited UTH1 and SKN7 yeast gene expression at 24 h. Amounts of the mRNA above were normalized to that of TUB1. The results were
displayed as mean ±SEM for three independent experiments (∗P<0 05,∗∗ P<0 01, and ∗∗∗P<0 001, compared with the control group).
7Oxidative Medicine and Cellular Longevity
group (Figure 4(c), P<001,P<001, and P<0 05), respec-
tively. These results suggested that compound 9 extended
the replicative lifespan via inhibition of oxidative stress.
3.5. Compound 9 Extends Yeast Lifespan via Modification of
UTH1, SKN7, SOD1, and SOD2 Gene Expression. It is well
known that antioxidative stress is one of mechanisms of
action for antiaging. UTH1 gene essentially takes part in
oxidative stress regulation, and deletion of UTH1 gene will
lead to extend the replicative lifespan of yeast [25]. SKN7 is
upstream gene and is a stress response transcription factor
in Saccharomyces cerevisiae [26].Superoxide dismutases
(SOD) are major ROS scavenging enzymes and can con-
vert superoxide anion to hydrogen peroxide [24]. Real-time
PCR analysis was performed to examine the molecular
mechanism of compound 9-mediated lifespan extension.
The significant gene expression reduction or increase of
UTH1,SKN7,SOD1, and SOD2 was observed in the com-
pound 9 treatment groups (Figure 5). These results suggested
that compound 9 produced antiaging effect via regulation
UTH1,SKN7,SOD1, and SOD2 yeast gene expression.
3.6. Antiaging Effects of Compound 9 Diminished in Uth1,
Skn7, Sod1, and Sod2 Mutations with K6001 Background.
To investigate the role of these genes in the antiaging activity
of compound 9, we used the mutants of uth1,skn7,sod1, and
sod2. As shown in Figure 6, compound 9 at 3 μM did not
affect the replicative lifespan of uth1 (Figure 6(a)) or skn7
mutants (Figure 6(b)), neither of sod1 (Figure 6(c)) or sod2
mutants (Figure 6(d)). These results were further indicated
that these four genes were involved in the mechanism of
action of compound 9.
4. Conclusions
A novel cucurbitane-type triterpenoid and nine known
compounds were isolated and identified from the fruits of
M. charantia. All the compounds showed antiaging effect in
0 2 4 6 8 10 12 14 16 18 20
0
20
40
60
80
100
Control (K6001)
Res at 10 M (K6001)⁎⁎⁎
9 at 3 M (K6001)⁎⁎
Control (Δuth1)
9 at 3 M (Δuth1)
Generations
Viability (%)
(a)
0 2 4 6 8 10 12 14 16 18 20
0
20
40
60
80
100
Control (K6001)
Res at 10 M (K6001)⁎⁎⁎
9 at 3 M (K6001)⁎⁎
Control (Δskn7)
9 at 3 M (Δskn7)
Generations
Viability (%)
(b)
024 6 810 12 14 16 18 20
0
20
40
60
80
100
Control (K6001)
Res at 10 M (K6001)⁎⁎⁎
9 at 3 M (K6001)⁎⁎
Control (Δsod1)
9 at 3 M (Δsod1)
Generations
Viability (%)
(c)
024 6 8 10 12 14 16 18 20
0
20
40
60
80
100
Control (K6001)
Res at 10 M (K6001)⁎⁎⁎
9 at 3 M (K6001)⁎⁎
Control (Δsod2)
9 at 3 M (Δsod2)
Generations
Viability (%)
(d)
Figure 6: Effect of compound 9 on the replicative lifespan of uth1 (a), skn7 (b), sod1 (c), and sod2 (d) mutants. The average lifespan of K6001
in the control group was 7.93 ±0.41; Res at 10 μM, 10.45 ±0.52∗∗∗; and compound 9 at 3 μM, 10.33 ±0.49∗∗. (a) The average lifespan of Δuth1
in the control group was 11.60 ±0.51 and compound 9 at 3 μM, 10.95 ±0.46. (b) The average lifespan of Δskn7 in the control group
was 9.88 ±0.41 and compound 9 at 3 μM, 10.40 ±0.53. (c) The average lifespan of Δsod1 in the control group was 6.55 ±0.32 and
compound 9 at 3 μM, 6.65 ±0.34. (d) The average lifespan of Δsod2 in the control group was 6.28 ±0.25 and compound 9 at 3 μM,
6.50 ±0.25 (∗∗P<0 01 and ∗∗∗P<0 001).
8 Oxidative Medicine and Cellular Longevity
yeast. The antiaging activities of these cucurbitane-type
triterpenoids depended on their antioxidative ability and
the regulation of the UTH1,SKN7,SOD1, and SOD2 yeast
genes. Apart from being one of the most well-known vegeta-
bles and frequently used as a traditional medicine because of
its health benefits, M. charantia has potential as an antiaging
functional food.
Conflicts of Interest
The authors declare no financial or commercial conflict
of interest.
Authors’Contributions
Xueli Cao and Yujuan Sun contributed equally to the article.
Acknowledgments
This work was supported by the NSFC (Grant nos.
21661140001, 81273385, and 21572204), International
Science and Technology Cooperation Program of China
(Grant no. 2014DFG32690), and Project of the Science and
Technology Department of Zhejiang Province (Grant no.
2015C33132). The authors are grateful to Akira Matsuura
(Chiba University, Japan) for the gifts of BY4741 and all
mutations with the K6001 background and grateful to
Michael Breitenbach (Salzburg University, Austria) for the
gifts of K6001.
References
[1] G. Nagarani, A. Abirami, and P. Siddhuraju, “Food prospects
and nutraceutical attributes of Momordica species: a potential
tropical bioresources –a review,”Food Science and Human
Wellness, vol. 3, no. 3-4, pp. 117–126, 2014.
[2] X. Xu, B. Shan, C. H. Liao, J. H. Xie, P. W. Wen, and
J. Y. Shi, “Anti-diabetic properties of Momordica charantia
L. polysaccharide in alloxan-induced diabetic mice,”Inter-
national Journal of Biological Macromolecules, vol. 81,
pp. 538–543, 2015.
[3] A. Raman and C. Lau, “Anti-diabetic properties and phyto-
chemistry of Momordica charantia L. (Cucurbitaceae),”Phyto-
medicine, vol. 2, no. 4, pp. 349–362, 1996.
[4] T. Murakami, A. Emoto, H. Matsuda, and M. Yoshikawa,
“Medicinal foodstuffs. XXI. Structures of new cucurbitane-
type triterpene glycosides, goyaglycosides-a, -b, -c, -d, -e, -f,
-g, and -h, and new oleanane-type triterpene saponins, goyasa-
ponins I, II, and III, from the fresh fruit of Japanese Momor-
dica charantia L.,”Chemical and Pharmaceutical Bulletin,
vol. 49, no. 1, pp. 54–63, 2001.
[5] J. K. Grover and S. P. Yadav, “Pharmacological actions and
potential uses of Momordica charantia: a review,”Journal of
Ethnopharmacology, vol. 93, no. 1, pp. 123–132, 2004.
[6] R. Horax, N. Hettiarachchy, and P. Chen, “Characteristics and
functionality enhancement by glycosylation of bitter melon
(Momordica charantia) seed protein,”Journal of Food Science,
vol. 79, no. 11, pp. C2215–C2221, 2014.
[7] Z.-J. Li, J.-C. Chen, Y.-Y. Deng et al., “Two new cucurbitane
triterpenoids from immature fruits of Momordica charantia,”
Helvetica Chimica Acta, vol. 98, no. 10, pp. 1456–1461, 2015.
[8] B. Zhang, C. Xie, Y. Wei, J. Li, and X. Yang, “Purification and
characterisation of an antifungal protein, MCha-Pr, from the
intercellular fluid of bitter gourd (Momordica charantia)
leaves,”Protein Expression and Purification,vol. 107, pp. 43–
49, 2015.
[9] O. Kenny, T. J. Smyth, C. M. Hewage, and N. P. Brunton,
“Antioxidant properties and quantitative UPLC-MS analysis
of phenolic compounds from extracts of fenugreek (Trigonella
foenum-graecum) seeds and bitter melon (Momordica char-
antia)fruit,”Food Chemistry, vol. 141, no. 4, pp. 4295–
4302, 2013.
[10] F. Zhang, L. Lin, and J. Xie, “A mini-review of chemical and
biological properties of polysaccharides from Momordica
charantia,”International Journal of Biological Macromole-
cules, vol. 92, pp. 246–253, 2016.
[11] M. Armanios, R. de Cabo, J. Mannick, L. Partridge, J. van
Deursen, and S. Villeda, “Translational strategies in aging
and age-related disease,”Nature Medicine, vol. 21, no. 12,
pp. 1395–1399, 2015.
[12] M. Kaeberlein, P. S. Rabinovitch, and G. M. Martin, “Healthy
aging: the ultimate preventative medicine,”Science, vol. 350,
no. 6265, pp. 1191–1193, 2015.
[13] Y. Weng, L. Xiang, A. Matsuura, Y. Zhang, Q. Huang, and
J. Qi, “Ganodermasides A and B, two novel anti-aging ergos-
terols from spores of a medicinal mushroom Ganoderma
lucidum on yeast via UTH1 gene,”Bioorganic & Medicinal
Chemistry, vol. 18, no. 3, pp. 999–1002, 2010.
[14] Y. Weng, J. Lu, L. Xiang et al., “Ganodermasides C and D, two
new anti-aging ergosterols from spores of the medicinal
mushroom Ganoderma lucidum,”Bioscience, Biotechnology,
and Biochemistry, vol. 75, no. 4, pp. 800–803, 2011.
[15] L. Xiang, K. Sun, J. Lu et al., “Anti-aging effects of phloridzin,
an apple polyphenol, on yeast via the SOD and Sir2 genes,”
Bioscience, Biotechnology, and Biochemistry, vol. 75, no. 5,
pp. 854–858, 2011.
[16] K. Sun, S. Cao, L. Pei, A. Matsuura, L. Xiang, and J. Qi,
“A steroidal saponin from Ophiopogon japonicus extends
the lifespan of yeast via the pathway involved in SOD and
UTH1,”International Journal of Molecular Sciences, vol. 14,
no. 3, pp. 4461–4475, 2013.
[17] Y. Lin, Y. Sun, Y. Weng, A. Matsuura, L. Xiang, and J. Qi,
“Parishin from Gastrodia elata extends the lifespan of yeast
via regulation of Sir2/Uth1/TOR signaling pathway,”Oxida-
tive Medicine and Cellular Longevity, vol. 2016, Article ID
4074690, 11 pages, 2016.
[18] T. Akihisa, N. Higo, H. Tokuda et al., “Cucurbitane-type
triterpenoids from the fruits of Momordica charantia and their
cancer chemopreventive effects,”Journal of Natural Products,
vol. 70, no. 8, pp. 1233–1239, 2007.
[19] H. Okabe, Y. Miyahara, and T. Yamauchi, “Studies on the
Constituents of Momordica charantia L. III. Characterization
of New Cucurbitacin Glycosides of the Immature Fruits.
Structures of Momordicosides G, F
1
,F
2
and I,”Chemical and
Pharmaceutical Bulletin, vol. 30, no. 11, pp. 3977–3986, 1982.
[20] S. Nakamura, T. Murakami, J. Nakamura, H. Kobayashi,
H. Matsuda, and M. Yoshikawa, “Structures of new
cucurbitane-type triterpenes and glycosides, karavilagenins
and karavilosides, from the dried fruit of Momordica charantia
L. in Sri Lanka,”Chemical & Pharmaceutical Bulletin, vol. 54,
no. 11, pp. 1545–1550, 2006.
[21] X. Wang, W. Sun, J. Cao, H. Qu, X. Bi, and Y. Zhao,
“Structures of new triterpenoids and cytotoxicity activities of
9Oxidative Medicine and Cellular Longevity
the isolated major compounds from the fruit of Momordica
charantia L.,”Journal of Agricultural and Food Chemistry,
vol. 60, no. 15, pp. 3927–3933, 2012.
[22] T. Tanaka, T. Nakashima, T. Ueda, K. Tomii, and I. Kouno,
“Facile discrimination of aldose enantiomers by reversed-
phase HPLC,”Chemical & Pharmaceutical Bulletin, vol. 55,
no. 6, pp. 899–901, 2007.
[23] F. Berrué, M. W. B. McCulloch, and P. Boland, “Isolation of
steroidal glycosides from the Caribbean sponge Pandaros
acanthifolium,”Journal of Natural Products, vol. 75, no. 12,
pp. 2094–2100, 2012.
[24] Y. Kong, S. E. Trabucco, and H. Zhang, “Oxidative stress,
mitochondrial dysfunction and the mitochondria theory
of aging,”Interdisciplinary Topics in Gerontology, vol. 39,
pp. 86–107, 2014.
[25] N. Camougrand, I. Kissova, G. Velours, and S. Manon,
“Uth1p: a yeast mitochondrial protein at the crossroads of
stress, degradation and cell death,”FEMS Yeast Research,
vol. 5, no. 2, pp. 133–140, 2004.
[26] V. Basso, S. Znaidi, V. Lagage et al., “The two-component
response regulator Skn7 belongs to a network of transcription
factors regulating morphogenesis in Candida albicans and
independently limits morphogenesis-induced ROS accumu-
lation,”Molecular Microbiology, vol. 106, no. 1, pp. 157–
182, 2017.
10 Oxidative Medicine and Cellular Longevity
Available via license: CC BY 4.0
Content may be subject to copyright.