Content uploaded by Eliana Regina Forni-Martins
Author content
All content in this area was uploaded by Eliana Regina Forni-Martins on Aug 06, 2021
Content may be subject to copyright.
Full Terms & Conditions of access and use can be found at
https://www.tandfonline.com/action/journalInformation?journalCode=tcar20
Caryologia
International Journal of Cytology, Cytosystematics and Cytogenetics
ISSN: 0008-7114 (Print) 2165-5391 (Online) Journal homepage: https://www.tandfonline.com/loi/tcar20
New chromosome counts in Neotropical
Machaerium Pers. species (Fabaceae) and their
taxonomic significance
Carlos Victor Mendonça Filho , Eliana R. Forni-Martins & Ana M.G.A. Tozzi
To cite this article: Carlos Victor Mendonça Filho , Eliana R. Forni-Martins & Ana M.G.A. Tozzi
(2002) New chromosome counts in Neotropical Machaerium Pers. species (Fabaceae) and their
taxonomic significance, Caryologia, 55:2, 111-114, DOI: 10.1080/00087114.2002.10589265
To link to this article: https://doi.org/10.1080/00087114.2002.10589265
Published online: 18 Dec 2012.
Submit your article to this journal
Article views: 172
View related articles
Citing articles: 1 View citing articles
INTRODUCTION
Widespread in the Neotropics, the genus
Machaerium comprises around 130 species, rang-
ing from Mexico to Argentina, many of which are
difficult to delimit taxonomically (RUDD 1986).
Most diversity occurs in southeastern Brazil and
in the Amazon basin. Traditionally, BENTHAM
(1860) divided this genus into five series, later
raised to sections by TAUBERT (1891). Closely re-
lated to Dalbergia, it was assigned, together with
12 other genera, to the Dalbergia group in the
tribe Dalbergieae, subfamily Faboideae (POLHILL
1994). However, recent cladistic analyses consid-
ered this tribe was not monophyletic and distin-
guished a Dalbergia clade that only included
Machaerium and Dalbergia, out of the former tribe
Dalbergieae, together with 14 genera from the
tribe Aeschynomeneae. They were then consid-
ered as more closely related to the Aeschynomene
species that have basifixed stipules (LAVIN et al.
2001).
This Dalbergia clade, together with other dal-
bergioid legumes and a group of Faboideae that
also includes the tribes Amorpheae and
Dipterygeae are fairly uniform cytologically and
have a base chromosome number x=10 (GOLD-
BLATT 1981; LAVI N et al. 2001).
The only chromosome numbers reported for
Machaerium concern M. aculeatum Raddi (n=10,
COLEMAN and DEMENEZES 1980), M. lunatum
(L.f.) Ducke (n=9, GILL and HUSAINI 1985a; HU-
SAINI and GILL 1986), from sect. Lineata; M. acu-
tifolium Vogel (2n=20, BANDEL 1974), from sect.
Acutifolia; and M. paraguariense (2n= 20, SAR-
CARYOLOGIA Vol. 55, no. 2: 111-114, 2002
New chromosome counts in Neotropical
Machaerium Pers. species (Fabaceae) and their
taxonomic significance
CARLOS VICTOR MENDONÇA FILHO, ELIANA R. FORNI-MARTINS* and ANA M.G.A. TOZZI
Departamento de Botânica, Instituto de Biologia, Universidade Estadual de Campinas, Caixa Postal 6109, Campinas
13083-970, SP, Brasil.
Abstract - Machaerium comprises about 130 species and is widespread in the
Neotropics, ranging from Mexico to Argentina. Because of the existing taxo-
nomic complexes, circumscribing many of its species is far from easy. Cytolog-
ical studies on Machaerium have only been carried out for four species and are
still meager. This paper brings forward chromosome numbers for 18 species, 16
of which represent new counts. Chromosomes were very small in size. Meiotic
behavior was observed only for M. opacum (n=10) and proved to be normal. The
other species showed 2n=20, except for the tetraploids M. nyctitans,M. hirtum
and M. sericiflorum, where 2n=40. Polyploidy possibly evolved independently
in Machaerium since these species belong to different sections. Both M. hirtum
and M. sericiflorum (tetraploids) were regarded as taxa distinct from M. aculea-
tum (diploid), although they used to be considered as synonyms. The impor-
tance of polyploidy for the evolution of species in this genus and its frequency
within the different sections will be assessed as more records become available.
It might also be possible to confirm whether its basic number is actually x=10,
as results tend to indicate.
Key words: chromosome numbers, cytotaxonomy, Fabaceae, Machaerium, poly-
ploidy.
* Corresponding author: fax ++ 55 19 32893124; e-mail:
elianafm@unicamp.br
TORI and FORNI-MARTINS, pers. com.), from sect.
Reticulata. Therefore, information is limited to
a small fraction of this genus, still poorly known,
cytologically speaking.
This study aims at extending the investigation
of chromosome numbers for this widespread
Neotropical genus to other species and sections,
trying to help circunscribing the taxa.
MATERIALS AND METHODS
Seeds of 17 Machaerium species and flower buds of
M. opacum, presenting different life forms and from
various habitats were collected in southern and north-
eastern Brazil. Most of them come from areas either of
forest or cerrado (a kind of savanna) (Table 1). Vouch-
ers were housed at the Herbarium (UEC) of the Botany
Department of the University of Campinas-SP. As for
meiotic studies, flower buds were fixed in 3:1
ethanol:acetic acid (Carnoy) and their anthers squashed
in acetocarmine (MEDINA and CONAGIN 1964). Young,
actively growing root-tips of each of the 17 species were
excised from potted plants and pretreated with satu-
rated solution of PDB (paradichlorobenzene) for 4
hours at 16-18ºC, in order to obtain a better chromo-
somal condensation and arrangement on the slide. The
material was fixed in Carnoy’s liquid for 24 h, and
stored in 70% ethanol. They were stained according to
the HCl/Giemsa method (GUERRA 1983). An Olympus
photomicroscope was used to photograph suitable
chromosome spreads. Permanent slides are kept at the
Biosystematic Laboratory of the Botany Department.
RESULTS
The 18 species of Machaerium studied belong
to four out of the five sections known for the
genus: sect. Reticulata (six species), sect. Acuti-
folia (six species), sect. Oblonga (three species),
and sect. Lineata (three species). The chromo-
somes were very small in size making it difficult
to determine karyotypes (Fig. 1, Table 1).
The chromosome analysis showed that all the
species studied presented 2n= 20, except for the
tetraploids M. nyctitans, M. hirtum and M. serici-
florum, with 2n=40 (Fig. 1, Table 1). The last two
species belong to sect. Lineata, and the first one
to sect. Oblonga. Meiotic behavior proved to be
normal in M. opacum and regular 10 bivalents
were observed at diakinesis (Table 1).
DISCUSSION
The chromosome counts found represent 16
new records for Machaerium. The numbers for
M. acutifolium Vogel and M. aculeatum Raddi
(2n=20) confirmed those found by BANDEL
(1974) and COLEMAN and DEMENEZES (1980),
respectively. This number was also detected for
M. paraguariense Vogel (SARTORI and FORNI-
MARTINS, pers. com.). Exception was M. lunatum
(L.f.) Ducke, with n=9 (GILL and HUSAINI 1985a;
HUSAINI and GILL 1986). The basic chromosome
number x=10 seems to fit Machaerium well and
agrees with records about both the tribe Dal-
bergieae and the dalbergioid legumes (GOLD-
BLATT 1981; LAVIN et al. 2001).
Polyploidy and the taxonomy of Machaerium
The polyploidy detected for M. nyctitans,
M. hirtum and M. sericiflorum is the first report
regarding Machaerium. As these species belong
to only two of the genus sections, sect. Oblonga
112 MENDONÇA FILHO, FORNI-MARTINS and TOZZI
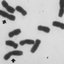
Fig. 1 – Chromosomes in Machaerium. A-B: tetraploid
species; C-F: diploid species. A) M. hirtum, B) M. serici-
florum, C) M. aculeatum, D) M. scleroxylon, E) M. villosum,
F) M. punctatum. Scale bar 10 µm.
AB
CD
EF
(M. nyctitans) and sect. Lineata (for the last two
species), it seems reasonable to suppose that poly-
ploidy has evolved independently within the
Machaerium sections and may furthermore be re-
stricted to these two sections of the genus. As for
Adesmia (Fabaceae, Faboideae, Adesmieae),
polyploidy could not be related to the serial in-
frageneric positions of species, which also indi-
cates its independent origin (MIOTTO and FORNI-
MARTINS 1994).
The assumption that polyploids are generally
widely distributed (GUERRA 1988) was support-
ed by the data on the first two species (very com-
mon species) but not by those on M. sericiflorum,
which presents a more restricted distribution.
Polyploidy played an important role in the
speciation of higher plants and all species of
Fabaceae cytologically known to date had a poly-
ploid ancestor (GOLDBLATT 1981; GILL and HU-
SAINI 1985b). However, polyploidy seems to play
a secondary role in the speciation of Machaerium
since it has only been recorded for three species,
so far.
The tetraploid number found for M. sericiflo-
rum and M. hirtum definitely separated them
from M. aculeatum (2n= 20). These species were
classified as M. aculeatum by HOEHNE (1941) and
many authors have followed his assumption, even
recently (LORENZI 1992). Although the floral
morphology of these species is similar, their fruit,
seed and seedling present markedly different
characters (MENDONÇA FILHO, in preparation).
Therefore, both karyotypic and morphological
characters give good evidence to consider these
species as distinct taxa.
A relationship between the species habits and
their classification into sections only seems possi-
ble for Machaerium sect. Acutifolia, where all
species are trees. As for the other sections, the
habit of the species ranges from lianas, shrubs up
to trees (Table 1). For the time being, no chro-
mosomal characters could be related to the dif-
ferences of habit observed within these sections.
Although it has been assumed that lianas derived
from trees (RADDFORD et al. 1974), and shrubby
plants occupy an intermediary position, trees be-
ing considered as less derived (CARLQUIST 1991),
the constancy in chromosome numbers (except
for the polyploids) and chromosome size observed
for the Machaerium species studied makes it im-
possible to set up any cytotaxonomic considera-
tions. LOMBELLO and FORNI-MARTINS (1998) sug-
gested that, as for Sapindaceae, the derivation of
lianas was followed by a chromosomal derivation,
through enlargement of the chromosomes and re-
duction of their number. In addition, with regard
to Malpighiaceae, another family with species
showing distinct habits, the tree species Lophan-
thera lactescens showed the lowest chromosome
number (2n= 12) and the longest chromosomes
(11.3 to 5.5 µm) (LOMBELLO and FORNI-MARTINS
1998).
CHROMOSOME NUMBERS IN MACHAERIUM 113
Table 1- Chromosome numbers and habits of the Machaerium species. The species were grouped into sections following BEN-
THAM (1860) except for those followed by (!), which have been rearranged according to MENDONÇA FILHO (in preparation).
Section/Species habit Voucher number (UEC) 2n/n* Ploidy level
sect. Reticulata
M. brasiliense Vogel Tree C.V. Mendonça 443 20 2x
M. lanceolatum (Vogel) J.F.Macbr. (!) Liana C.V. Mendonça 439 20 2x
M. mucronulatum Mart. ex Benth. (!) Shrub C.V. Mendonça 658 20 2x
M. oblongifolium Vogel Liana T.S. Spósito s.n. 20 2x
M. opacum Vogel Tree C.V. Mendonça 533 10* 2x
M. triste Vogel Liana C.V. Mendonça 547 20 2x
sect. Acutifolia
M. acutifolium Vogel Tree C.V. Mendonça 446 20 2x
M. fulvovenosum H.C. Lima (!) Tree C.V. Mendonça 600 20 2x
M. pedicelatum Vogel (!) Tree E.T. Neto s.n. 20 2x
M. punctatum Pers. (!) Tree C.V. Mendonça 548 20 2x
M. stipitatum Vogel Tree C.V. Mendonça 450 20 2x
M. villosum Vogel Tree C.V. Mendonça 434 20 2x
sect. Oblonga
M. nyctitans (Vell.) Mart. ex Benth. Tree C.V. Mendonça 428 40 4x
M. scleroxylon Tul. Tree C.V. Mendonça 549 20 2x
M. uncinatum Benth. (!) Liana C.V. Mendonça 441 20 2x
sect. Lineata
M. aculeatum Raddi Liana C.V. Mendonça 606 20 2x
M. hirtum (Vell.) Stellfeld Tree C.V. Mendonça 437 40 4x
M. sericiflorum Vogel (!) Liana C.V. Mendonça 633 40 4x
Chromosomal data only exist for a small frac-
tion (14%) of the about 130 species of Machaeri-
um. This study must be extended to other
species in order to verify the existence of a pos-
sible relationship between the derivation of kary-
otypic characters and changes of morphological
characters.
The cytological knowledge must be used in
conjunction with other sources of data, from as
many different areas as possible, so as to achieve
a better understanding of the phylogenetic rela-
tionship of species in the genus and come the best
classification.
Acknowledgments – The authors would partic-
ularly like to thanks Yara Bressan for laboratory as-
sistance and Teresa Sposito for providing seeds of
M. oblongifolium as well as the Fundação de Am-
paro à Pesquisa do Estado de São Paulo for the
grants 98/02889-3 and 98/6378-3.
REFERENCES
BANDEL G., 1974 – Chromosome numbers and evo-
lution in the Leguminosae. Caryologia, 27: 17-
32.
BENTHAM G., 1860 – Synopsis of Dalbergieae a tribe
of Leguminosae. Journal of the Linnean Society,
Botany, 4: 1-128.
CARLQUIST S. 1991 – Anatomy of vine and liana
stems: a review and synthesis. In F.E. Putz and
H.A. Mooney (Eds), “The Biology of Vines”,
pp. 53-72. University of Cambridge Press, New
York.
COLEMAN J.R. and DEMENEZES E.M. 1980 – Chro-
mosome numbers in Leguminosae from the State
of São Paulo, Brazil. Rhodora, 82: 475-481.
GILL L.S. and HUSAINI S.W.H., 1985a – Cytology of
some arborescent Papilionoideae (Leguminosae)
of southern Nigeria. Boletim Sociedade Broteri-
ana, 2: 187-200.
–, 1985b – Caryological evolution of southern Niger-
ian Leguminosae. Revue de Cytologie et de Bi-
ologie Végetales, 8: 3-31.
GOLDBLATT P., 1981 – Cytology and Phylogeny of
Leguminosae. In R.M. Polhill and P.H. Raven
(Eds), “Advances in Legume Systematics”, part.
2, pp. 427-463, Royal Botanic Gardens, Kew.
GUERRA M.S., 1983 – O uso do corante Giemsa na
citogenética vegetal- comparação simples e o ban-
deamento. Ciência e Cultura, 35: 190-193.
–, 1988 – Introdução a Citogenética Geral. Guan-
abara S.A., Rio de Janeiro.
HOEHNE F.C., 1941 – Leguminosas Papilionadas
(Machaerium e Paramachaerium). Flora Brasili-
ca, 25: 1-99.
HUSAINI S.W.H. and GILL L.S., 1986 – Cytology of
the tribe Dalbergieae (Leguminosae) from Nige-
ria. Feddes Repertorium, 97: 469-473.
LAVIN M., PENNINGTON R.T., KLITGAARD B.B.,
SPRENT J.I., LIMA H.C. and GASSON P.E., 2001
– The Dalbergioid Legumes (Fabaceae): delimi-
tation of a pantropical monophyletic clade. Amer.
J. Bot., 88: 503-533.
LOMBELLO R.A. and FORNI-MARTINS E.R., 1998 –
Chromosomal studies and evolution in Sapin-
daceae. Caryologia, 51: 81-93.
LORENZI H., 1992 – Árvores Brasileiras: manual de
identificação e cultivo de plantas arbóreas nativas
do Brasil. Plantarum, Nova Odessa.
MEDINA D.M. and CONAGIN C.H.T.M., 1964 – Téc-
nica citológica.Publicação 2610. Instituto
Agronômico, Campinas.
MIOTTO S.T.S. and FORNI-MARTINS E.R., 1994 –
Número cromossômico em espécies brasileiras de
Adesmia DC. (Leguminosae-Faboideae). Acta
Botânica Brasilica, 8: 3-9.
POLHILL R.M., 1994 – Classification of the Legumi-
nosae. In F.A. Bisby, J. Buckingham and J.B.
Harborne (Eds), “Phytochemical dictionary of
the Leguminosae”, p. 11. Chapman and Hall,
New York.
RADFORD A.E., DICKINSON W.C., MASSEY J.R. and
BELL C.R., 1974 – Vascular Plant Systematics.
Harper and Row Publ., New York.
RUDD V.E., 1986 – A new species of Machaerium
(Leguminosae) from Nicaragua. Phytologia, 60:
93-94.
TAUBERT P., 1891 – Machaerium: In H.G.A. Engler
and K.A.E. Plantl (Eds), “Die Naturlichen
Pflanzenfamilien”, vol. 3, Leipzig.
Received October 24, 2001; accepted December 5, 2001
114 MENDONÇA FILHO, FORNI-MARTINS and TOZZI