Content uploaded by Oussama Abchir
Author content
All content in this area was uploaded by Oussama Abchir on Sep 24, 2022
Content may be subject to copyright.
Vol.:(0123456789)
1 3
Journal of Molecular Modeling (2022) 28:106
https://doi.org/10.1007/s00894-022-05097-9
ORIGINAL PAPER
Design ofnovel benzimidazole derivatives aspotential α‑amylase
inhibitors using QSAR, pharmacokinetics, molecular docking,
andmolecular dynamics simulation studies
OussamaAbchir1· OssamaDaoui2· SalahBelaidi3· MebarkaOuassaf3· FaizanAbulQais4· SouadElKhattabi2·
SaidBelaaouad1· SamirChtita5
Received: 8 September 2021 / Accepted: 15 March 2022
© The Author(s), under exclusive licence to Springer-Verlag GmbH Germany, part of Springer Nature 2022
Abstract
In the present study, a quantitative relationship between the biological inhibitory activity of alpha-amylase and molecular struc-
tures of novel benzimidazole derivatives is analyzed in silico. The best QSAR model screened via MLR technique indicated
that the exact mass, topological diameter and numerical rotational bonding structural properties of benzimidazole derivatives
highly affect the bioactivity of these compounds against α-amylase. Based on the structural properties identified via linear QSAR
model favorable for improving pIC50 of benzimidazole derivatives, fourteen new molecules bearing benzimidazole radicals
were designed and their biological inhibitory activity against α-amylase was improved. QSAR model predictions showed that the
designed molecules exhibited a higher potential biological level activity IC50 than acarbose used in positive control (IC50= 1.46
μM). Screening of drug-like properties, pharmacokinetics and toxicity of the proposed molecules led to select three molecules
as candidates for use as drug aid to ingest starch and glycogen. As a result, using molecular docking simulations, the docking
poses of the three molecules inside the α-amylase receptor pocket (PDB code: 1HNY) were predicted. Also, the most important
potential interactions between the active amino acid sites in α-amylase protein pocket and the proposed drug molecules were
described. The obtained hypotheses regarding the stability of the proposed molecules inside α-amylase pocket were validated
by carrying out molecular dynamic simulations in aqueous background similar to the ones of proteins. The DM results con-
firmed the optimal stability of the α-amylase backbone with the drug molecules proposed in this computational investigation.
Keywords QSAR· ADMET· Molecular docking· Molecular dynamics· Alpha-amylase· Benzimidazole
Introduction
Diabetes mellitus (DM) is one of the important causes of mor-
bidity and mortality. It was estimated in 2011 that 347 million
people were affected by DM worldwide, and it will get dou-
bled in 2030. Diabetes mellitus can be defined by a type of
metabolic disorder when the patient has an increase in levels
of sugar (glucose) in the blood [1], which can lead to poten-
tial complications that include blindness, heart attacks, stroke,
kidney damage, lower limb amputation, and nerve damage [2].
The most common type of diabetes is called type 2 diabe-
tes (formerly called non-insulin-dependent, or adult onset)
[3–5]. It happens when the pancreas stops using insulin
effectively, which means the body’s cells lose the ability to
respond to insulin’s efforts to drive glucose into the cells, a
condition called insulin resistance [6].
As treatment of diabetes mellitus, many different strategies
were used, which are increasing the action of insulin, reduction of
* Samir Chtita
samirchtita@gmail.com
1 Laboratory ofPhysical Chemistry ofMaterials, Faculty
ofSciences Ben M’Sik, Hassan II University ofCasablanca,
Casablanca, Morocco
2 Laboratory ofEngineering, Systems andApplications,
National School ofApplied Sciences, Sidi Mohamed Ben
Abdellah-Fez University, BP72Fez, Morocco
3 Group ofComputational andMedicinal Chemistry, LMCE
Laboratory, University ofBiskra, BP 145 , 707000Biskra,
Algeria
4 Department ofAgricultural Microbiology, Faculty
ofAgricultural Sciences, Aligarh Muslim University,
AligarhUP-202002, India
5 Laboratory ofAnalytical andMolecular Chemistry, Faculty
ofSciences Ben M’Sik, Hassan II University ofCasablanca,
Sidi Othman, Box7955, Casablanca, Morocco
Journal of Molecular Modeling (2022) 28:106
1 3
106 Page 2 of 17
the demand for insulin, encouraging the endogenous secretion of
insulin, or inhibiting the degradation of oligo and disaccharides.
For the purpose to lower blood glucose levels, it is needed
to inhibit alpha-amylase activity, which is a metalloenzyme
adopted Ca2+ ion on its active site that boosts the hydrolysis of
starch into glucose and maltose [7]. The high quantity of car-
bohydrate uptake leads to severe health such as diabetes [8].
Currently, acarbose is used as a medication for inhib-
iting the alpha-amylase [9], but it causes many adverse
effects such as diarrhea, abdominal discomfort [10]. For
this, researchers try to find new therapies for the manage-
ment of type 2 diabetes mellitus with no or low side effects.
Many studies have been done to treat this disease using
benzimidazole derivatives that can be defined by an aro-
matic heterocyclic organic compound having a benzene
ring fused with an imidazole ring as shown in Table1.
Imidazole ring is part of many natural products including
purine, histamine, histidine, and nucleic acid. Due to its
polar and ionizable ability, it proves to be characteristic
pharmacokinetics for lead molecules by enhancing their
solubility. Benzimidazole is very used in medicinal chemis-
try due to its activities including antimicrobial, anticancer,
antiviral, and antidiabetic activities.
Recently, molecular modeling techniques have emerged
as a promising approach to design and develop novel com-
pounds with interesting biological properties [11, 12]. In
this regard, we used in the current work QSAR studies
based on the assumption that the activity of chemical com-
pounds related to its structure through a certain mathemati-
cal algorithm. This relationship can be used in the predic-
tion, interpretation, and assessment of new compounds with
desired activities reducing and rationalizing time, efforts
and cost of synthesis, and new product development. We
also perform molecular docking and molecular dynamics
(MD) techniques to predict potential new molecules and
the active amino acid residue sites in the alpha-amylase
target, and consequently to evaluate the potential interac-
tions between these of the designed candidate molecules as
promising agents against the alpha-amylase enzyme.
In a previous study [13], Adegboy etal. synthesized and
analyzed a series of 45 benzimidazole derivatives to determine
their alpha-amylase inhibitory concentrations and compared
them to acarbose used as a reference. The researchers used
SAR analysis to identify the chemical group responsible for
inhibiting alpha-amylase activity, followed by a molecular
docking study to support their findings by studying the binding
mechanism of 2-aryl benzimidazole derivatives in the active
site of alpha-amylase. However, the results of this study sug-
gest that the synthesized compounds have a higher value of
IC50 than acarbose.
In the present work, we have built QSAR models using
multiple linear regression as a linear technique to design new
drug candidates with good α-amylase inhibitory activity greater
than the acarbose. In addition, we have used drug-likeness and
pharmacokinetic properties to select the “hits” that are suitable
starting points for research on a new clinical candidate. Then,
we have used molecular docking to study the interactions of
selected hits in the active site of alpha-amylase and we analyzed
the stability of various ligands poses under MD simulation.
Material andmethods
A QSAR study has been applied on a series of 37 benzimi-
dazole derivatives with their experimental activity values
IC50 (the half-maximal inhibitory concentration) expressed
in µmol that were compiled from the previous work [13].
The molecular structures of the studied molecules with their
activity are presented in Table1. To form a dataset, molecu-
lar descriptors of these molecules were calculated by using
Chemoffice, Marvinsketch, and Chemsketch [14–16].
Principal components analysis (PCA)
The PCA method available in XLSTAT software [17] was
applied to predict the correlation between molecular descrip-
tors with their alpha-amylase activities with the purpose to
reduce the size of the data representation space without los-
ing the information containing efficient, simple, and under-
standable data [18, 19].
Multiple linear regressions (MLR)
The stepwise regression linear multiple (MLR) method avail-
able in XLSTAT software was used for the purpose to deter-
mine the physicochemical effects of the substituents on the
activities of molecules. It was employed to find a linear model
of the studied activity, which takes the form below [20, 21]:
where Y: the studied activity, which is, the dependent vari-
able.
a0
: the intercept of the equation.
xi
: the molecular
descriptors.
ai
: The coefficients of those descriptors.
This method is one of the most popular methods of QSAR
thanks to its simplicity in operation, reproducibility, and
ability to allow easy interpretation of the features used. The
important advantage of the linear regression analysis is that
it is highly transparent; therefore, the algorithm is available
and predictions can be made easily.
After splitting the dataset into a training set consisting
of 30 compounds and a test set as a role in a 1/5 ratio [22],
MLR models were constructed on the training-set com-
pounds and the test-set compounds were used to predict
externally the resulting model.
Y
=a0+
n
∑
i=1
aix
i
Journal of Molecular Modeling (2022) 28:106
1 3
Page 3 of 17 106
Table 1 Chemical structures of studied benzimidazole derivatives with its observed activities IC50
NH
N
R
3
R
1
R
2
N° Structures IC50
(µM) N° StructureIC50
(µM) N° StructureIC50
(µM)
R1=R2=H
1 2.74
R1=R2=CH3
5 2.99 10 1.48
2 2.48 6 2.78 11 1.59
3 1.51 7 2.84 12 1.77
4 2.85 8 2.3 13 2.92
9 2.26 14 2.98
R1=H R2=F
15 2.98 19 1.68 23 1.57
16 1.59 20 2.54 24 2.56
17 2.60 21 2.49 25 2.87
18 2.62 22 2.17 26 2.86
R1=H R2=Cl
27 1.68 28 2.78 29 2.62
30 2.74 31 1.73 32 1.61
33 2.86 34 1.51 35 1.62
36 2.21 37 2.52
Journal of Molecular Modeling (2022) 28:106
1 3
106 Page 4 of 17
Model validation
The proposed models were validated by an internal valida-
tion method using leave-one-out cross-validation (LOOCV)
to characterize robustness and then by an internal validation
to estimate the predictive power of the models. To select the
best MLR model, coefficient of determination (R2), coefficient
adjusted for degrees of freedom (R2adj), coefficient of determi-
nation for the test set (R2test), and cross-validation coefficient
(R2CV) were used; R2, R2adj, R2test, and R2CV values greater than
0.6 indicates that the model is acceptable [23, 24].
Y-Randomization test was used to ensure the robust-
ness of the predictive models by comparing R2 of the built
model for permuted response and the original models
building procedure [25, 26].
Finally, the applicability domain was used to validate
the suggested models by using William’s plot available
in Matlab [27, 28]. It was used to define areas where it
is possible to practically use the model with increased
confidence about the prediction so obtained. The domain
of applicability is established in a squared area within a
standard deviation (x = 3) used as a cutoff value for accept-
ing predictions [29, 30].
Designing ofnew molecules
All studies allow choosing the best model that will be used
to design new molecules with good predicted activity val-
ues against alpha-amylase.
The leverage values hi of designed molecules were calcu-
lated (hi = xit × (Xt × X)−1 × xi (i = 1, 2, 3…n)) and compared
with the warning leverage h* (h* = 3*(k − 1)/n). The leverage
(hi) less than the warning leverage (h*) suggested that the
compound belongs to the applicability domain of the model.
Where: xi: the matrix of model descriptors of each designed
molecule, X: the matrix of model descriptor values for n train-
ing set compounds, the superscript t refers to the transpose of
matrix/vector of designed molecules, n is the number of train-
ing set compounds, and k is the number of model descriptors.
Pharmacokinetic profile
We further predicted the drug-like behavior of the compounds
through the analysis of pharmacokinetic parameters, of the
compounds by using the SwissADME web tool [31, 32].
Swiss ADME tool has been used to predict the
passive human gastrointestinal absorption (GI), pen-
etration of the blood–brain barrier (BBBP), skin pen-
etration, and inhibition of the human cytochromes
(CYPs): CYP1A2, CYP2C9, CYP2C19, CYP2D6, and
CYP3A4.
The toxicity evaluation was performed also using an
online server (ProTox-II) that gives predicted toxicity
values, cytotoxicity, mutagenicity, carcinogenicity, and
immunotoxicity [33–35].
Molecular docking
Molecular docking studies have been performed to deter-
mine the binding affinity and predict the molecular interac-
tions implicated between the active sites of the target protein
and some ligands [36]. The molecular docking method has
been used to complete this study by determining the best
poses of proposed molecules and that of acarbose at the
active site of the alpha-amylase for the purpose of compar-
ing them. The negative and low value of binding energy
obtained demonstrates a favorable conformation between
ligands and protein selected [37].
The first step was to download the alpha-amylase struc-
ture file (PDB code: 1HNY) [38] from the protein data bank
(www. rcsb. org). And the structures of ligands were drawn
and optimized by using Sybyl software [39].
Using Autodock tools [40], water and small molecules
co-crystallized were removed from the protein structures,
and polar hydrogens and Kollmann charges were added to
the structures in the aim of the prepare the protein before
starting the docking [38]. A grid box with a size of x = 86,
y = 74, and z = 76, and a center of x = 8.581, y = 60.249, and
z = 21.229 were set to cover the binding site of protein. The
AutoDock Vina software [41] was used to achieve the dock-
ing analysis to evaluate the ligand binding and interactions
with the alpha-amylase protein. The 2D and 3D interactions
of the docking results were obtained by importing our result
to the Discovery Studio Visualizer, hence, enabling us to
identify significant interactions between the ligands and the
enzyme binding site [42].
Molecular dynamics simulations
The top two compounds (A2 and A13) exhibiting the
lowest binding energy in molecular docking were
taken further for molecular simulation and dynam-
ics studies. The molecular dynamics (MD) simulation
studies were conducted using GROMACS-2018.1
packages with amber99sb-ILDN force field [43, 44].
The topology of ligands was generated in antechamber
packages in AmberTools19 [45]. The α-amylase alone
and their complexes with A2 and A13 were solvated
in a triclinic box separately using the TIP3P water
model. The structures were neutralized by adding
counter ions. To get rid of weak van der Waals con-
tacts, the systems were minimized for 5000 steps using
the steepest descent minimization. Following energy
minimization, the systems were equilibrated for NVT
Journal of Molecular Modeling (2022) 28:106
1 3
Page 5 of 17 106
and NPT each for 1ns using a V-rescale thermostat
at 300K and using Parrinello–Rahman barostat at
1.0bar [46, 47]. Standard 100ns MD simulation was
performed for each system, and 10,000 frames were
recorded in each trajectory at 10ps intervals. PBC
(periodic boundary conditions) corrections in each tra-
jectory were done before analysis. The analysis of tra-
jectory was performed using GROMACS utilities. For
the determination of various binding energies, MM-
PBSA calculation was done by extracting 50 frames
from 50 to 100ns of the trajectory [48].
Results anddiscussion
QSAR studies
We formed the data set by calculating 72 molecular
descriptors for the 37 benzimidazole derivatives, and we
used principal component analysis to eliminate descrip-
tors that correlated each with the other without losing the
initial information. We reduced the number of descrip-
tors from 72 to 35 (critical pressure, boiling point, heat
of formation, critical temperature, Henry’s law constant,
melting point, partition coefficient, Connolly accessible
area, exact mass, number of H bond acceptors, percent
composition of compounds (C, H, N, and O), number of H
bond donors, num rotatable bonds, Balaban index, polar
surface area, radius atom, shape attribute, shape coeffi-
cient, sum of valence, topological diameter, total valence
connectivity, index of refraction, parachor, surface ten-
sion, density, atom count, hydrophilic-lipophilic balance,
dreiding energy, min projection area, max projection area,
min projection radius, and max projection radius). We
applied a stepwise strategy to develop MLR models that
we got more than 20 models internally validated by cal-
culating R2, R2adj, and externally by randomization and
calculating Q2 [49, 50]. The majority of the molecular
models obtained show that the activity of alpha-amylase
is influenced by the exact mass, topological diameter, and
num rotatable bonds. As a result, we choose the model
number 1 (Eq.1) that have a good value of (R2 = 0.674,
R2adj = 0.650, R2test = 0.622, Q2 = 0.601).
For our 37 compounds, the correlation between
experimental activity and calculated one based on this
model is quite significant (Fig.1) that it shows a very
regular distribution of activity values depending on the
experimental values.
Y‑randomization test
In the present case, the results of 100 random shuffles for
the Y-randomization test show that none of the random tri-
als could match the original model as shown in TableS1.
The Q2 and R2 values of the new QSAR models were really
low. Therefore, the possibility of random correlations was
ruled out.
The average values of R, R2, and R2CV are 0.253, 0.081,
and − 0.137, respectively, the cRp2 value equals 0.640 (more
than 0.5), and all the new MLR models having significantly
low R2 and R2CV values for the 100 trials, which confirm that
the developed QSAR models are robust (TableS1).
(1)
IC50 =8087 −0.007exactmass − 0.341topologicaldiameter
Ttest (exactmass)=−5.001;Ttest (t opologicaldiameter )=−5.557
Fig. 1 Correlations of observed
and predicted activities (training
set in blue and test set in red)
values calculated using MLR
models
1.4
1.6
1.8
2
2.2
2.4
2.6
2.8
3
3.2
1.4 1.6 1.8 22.2 2.42.6 2.833.2
IC50
Pred(IC50)
Pred(IC50) / IC50
Acves Validaon
Journal of Molecular Modeling (2022) 28:106
1 3
106 Page 6 of 17
Domain ofapplicability
The applicability domain (AD) of this model was evaluated
by leverage analysis expressed as Williams plot (Fig.2), in
which the standardized residuals (r) and the leverage thresh-
old values (h ∗ = 0.250) were plotted. Any new value of pre-
dicted IC50 data must be considered reliable only for those
compounds that fall within this AD on which the model was
constructed. The results showed that there is no response
outlier in the training set and no response outside of the test
set. All the compounds have a standard deviation into ± x
interval (x = 3), and their leverages are inferior to leverage
threshold h* = 0.250.
Design ofnovel compounds
The IC50 value against a-amylase must be smaller than
acarbose (IC50 = 1.48). The results of a-amylase inhibi-
tion assay applied on the studied compound by Adegboy
etal. indicated that among the studied compounds, only
compounds 3, 10, 11, 16, 23, and 34 showed an activ-
ity similar to acarbose, and the common link between
these compounds is the presence of a methoxy group
or halogens elements. As well as the molecular model
developed by using RLM helps us to understand, which
molecular descriptors impact alpha-amylase activity.
According to these results, we tried to increase the val-
ues of exact mass, number rotatable bonds, and topo-
logical diameter by adding other methoxy groups and
halogens for the purpose to reduce the value of alpha-
amylase activity.
Based on developed model descriptors we have proposed
14 new molecules with an IC50 inferior to that of the acar-
bose, increasing the values of exact mass and topological
diameter in the designed molecules decrease the values of
their IC50.
The presence of a methoxy group increases the
inhibitory potential in potent derivatives. Wherever,
the absence of a methoxy group and the presence of
hydroxyl, bromo, and chloro in the benzimidazole
ring decreased the inhibitory effect. It appears that the
hydroxy group, Bromo, and chloreare not participating
in the inhibitory potential.
We calculated the leverages of all molecules designed
compared with the leverage threshold h*. The leverages of
designed molecules must be inferior to (h* = 0.243), and the
results shown in Table2 provide that all thus molecules are
acceptable except for compound A4 (h4 = 0.318).
Pharmacokinetic, physicochemical parameters,
anddrug‑likeliness studies
Drug-likeness and pharmacokinetic properties prediction of
the 14 designed compounds were performed by the online
version of SwissADME, and the data is shown in Table3.
The prediction of drug-likeness was also made based on
the selected Lipinski, Ghose, and Veber rules and the bio-
availability score [51–53].
Fig. 2 Williams plot of
standardized residual versus
leverage for the MLR model
(with h* = 0.250 and residual
limits ± 3); train samples in
black color and test samples in
red color)
Journal of Molecular Modeling (2022) 28:106
1 3
Page 7 of 17 106
Table 2 Designed molecules
with values of their molecular
descriptors, leverages hi, and
IC50
NIC50 Structure Topological
Diameter
Exact
mass hi
A1 1.307 11 411.058 0.080
A2 1.379 12 355.189 0.043
A3 1.404 11 398.026 0.063
A4 1.170 10 475.802 0.318
A5 1.183 11 427.902 0.108
A6 1.426 11 394.976 0.060
A7 1.315 11 410.012 0.079
A8 1.462 11 390.059 0.054
A9 0.464 13 433.028 0.052
A10 1.300 11 412.003 0.082
Journal of Molecular Modeling (2022) 28:106
1 3
106 Page 8 of 17
The Lipinski, Ghose, and Veber rules were applied to
assess drug-likeness to predict whether a compound is likely
to be bioactive according to some important parameters such
as molecular weight, LogP, number of HBA, and HBD.
The screening process with Lipinski rule of five Veber, and
Table 2 (continued)
A11 1.418 11 396.047 0.061
A12 0.947 11 459.942 0.179
A13 1.463 10 436.064 0.193
A14 1.389 11 399.983 0.066
Table 3 Drug-likeness prediction for the proposed compounds by SwissADME
MW g/mol LogP NR HBA HBD REF TPSA (Å2) Solubility Bioavail-
ability score Pains alert Synthetic
accessibility
score
A1 412.28 2.91 2 4 2 107.94 71.03 Moderately 0.55 0 2.88
A2 355.43 1.36 4 4 1 103.95 59.61 Soluble 0.55 0 5.03
A3 399.24 2.96 3 4 2 101.74 67.37 Moderately 0.55 0 2.43
A4 477.92 4.86 1 1 1 89.65 28.68 Poorly 0.55 0 2.40
A5 429.05 3.87 2 2 1 88.44 37.91 Moderately 0.55 0 2.35
A6 395.15 2.22 3 4 1 89.56 83.73 Moderately 0.55 0 2.72
A7 410.21 2.57 4 4 1 93.72 56.37 Moderately 0.55 0 2.57
A8 390.26 4.79 3 1 1 98.76 28.68 Poorly 0.55 0 2.76
A9 433.24 3.02 3 3 1 106.64 50.38 Moderately 0.55 0 3.67
A10 412.18 1.70 4 6 1 89.31 82.15 Moderately 0.55 0 2.95
A11 397.27 3.51 3 3 1 103.35 47.14 Moderately 0.55 0 3.25
A12 462.13 4.12 3 3 1 107.42 47.14 Poorly 0.55 0 2.53
A13 450.31 4.03 3 3 0 112 36.28 Poorly 0.55 0 3.00
A14 398.63 3.98 2 2 1 90.71 37.91 Moderately 0.55 0 2.42
Journal of Molecular Modeling (2022) 28:106
1 3
Page 9 of 17 106
Ghose showed that all compounds meet the criteria of drug-
likeness assessment (Table3).
The prediction of bioavailability (Table4) revealed
that the same bioavailability scores were obtained for all
studied molecules (0.55). This criterion is based on the
probability value of a compound to possess an optimum
profile of bioavailability and permeability, where the
value of 0.55 implies the obedience of the Lipinski rule
of five and 55% probability of rat bioavailability value
higher than 10%.
Molecules targeting oral administration, solubility is one
foremost property that influences absorption to deliver an
adequate quantity of active ingredients in a small volume,
Table4 suggests that the compounds A1, A2, A3, A5, A6,
A7, A9, A10, A11, and A14 have moderate water solubility,
and thus could facilitate good oral adsorption.
PAINS are important features to be considered while
developing drugs to avoid false positives results. In pan-
assay interference compounds (PAINS), structural alert
obtained 0 violation for all compounds.
Synthetic accessibility score shows compounds propose
in this study are easy synthetically (between 2.19 and − 3.14)
(Table3).
The ADME properties of the selected compounds
such as HIA rate and BBB penetration the predicted
values of these properties are shown in Table6. The
GI absorption for previous compounds is high; the
blood–brain barrier (BBB) permeation expresses the
relative affinity of the drug for the blood or brain tis-
sue. The blood–brain permeability was observed for
all compounds except A1, A6, A10, and A12. Com-
pounds A1, A6, A10, and A12 were observed perme-
ability negative. In the case of skin permeation (log
Kp, cm/s), a higher negative value was obtained for all
compounds < − 4.
P-gp is responsible for efflux across biological mem-
branes of a wide range of therapeutic drugs. One major
role of P-gp is to protect the central nervous system (CNS)
from xenobiotics [54]. It should be noted that compounds
A1, A2, and A12 have a high probability of being a sub-
strate of P-gp.
The cytochrome P450 (CYP) superfamily is important
in the elimination of drugs by metabolic biotransformation.
The inhibition of these isoenzymes is certainly a major cause
of drug interactions linked to pharmacokinetics [55]. The
result showed that the compounds A2, A4, A5, A6, A10,
A11, and A13 interact with at least one of the five isoforms,
while the compounds A1, A3, A7, A8, A9, and A12 are
found to be inhibitors activity for cytochrome P450, so there
may be a side effect (i.e., liver impairment).
The in silico toxicity prediction using a ProTox-II web
server showed that, except A1, A3, A7, A8, A9, and A12,
all other compounds were able to pass the ADME profile
to the toxicity side study, and except A4, all compounds
are found to have mutagenicity. Compounds A5, A6, and
A14 are reported to be carcinogenic. Compounds A2, A10,
and A13 toxicity levels were found to be within permissible
limits for properties such as carcinogenic, immunotoxicity,
cytotoxic, and hepatotoxicity models.
Compound with higher values of the predicted median
lethal dose (LD50) presents a lower toxic effect. Compounds
A10, A11, and A13 compounds have an LD50 of ˃600mg/
kg BW, A2 has an LD50 of 120mg/kgBW, while A4, A5,
A6, and A14 have an LD50 of < 100mg/kgBW, so A4, A5,
A6, and A14 were not selected for the lowest toxic effect
(Table5).
Table 4 Pharmacokinetics prediction for the proposed compounds by SwissADME
GI absorption BBB permeant P-gp substrate Cyp1A2 Cyp2c19 Cyp2c9 Cyp2d6 Cyp3a4 Log kp
A1 High No Yes Yes Yes Yes Yes Ye s − 5.41
A2 High Yes Yes No No No Yes No − 6.63
A3 High Yes No Yes Yes Yes Yes Yes − 5.41
A4 High Yes No Yes No Yes No No − 5.54
A5 High Yes No Yes Yes Yes No Yes − 5.75
A6 High No No Yes Yes Yes No No − 6.15
A7 High Yes No Yes Yes Yes Yes Yes − 6.17
A8 High Yes No Yes Yes Yes Yes Yes − 4.59
A9 High Yes No Yes Yes Yes Yes Yes − 6.71
A10 High No No Yes No Yes Yes Yes − 6.69
A11 High Yes No Yes Yes Yes No Yes − 5.51
A12 High No Yes Yes Yes Yes Yes Ye s − 5.06
A13 High Yes No No Yes Yes No No − 5.39
A14 High Yes No Yes Yes Yes No Yes − 5.35
Journal of Molecular Modeling (2022) 28:106
1 3
106 Page 10 of 17
Molecular docking
The results of molecular docking were obtained by
importing our result to the discovery studio visualizer,
and it shows the binding affinity among the 4 docked
ligands (three screened by the ADMET study and the
acarbose as reference) into the binding site of the 1HNY
as shown in Table6.
The 2D and 3D viewing of the protein–ligand interac-
tions of the best poses engender by the two ligands studied
and that of acarbose is illustrated in Fig.3.
Compounds A2, A13, and acarbose showed the best
binding score with the protein 1HNY are mentioned in
Table6.
Binding analysis within the active site of alpha-amylase
revealed that the acarbose forms conventional hydrogen
bond and carbon-hydrogen bond with the amino acid resi-
due of ASP402, TRP280, HIS331, GLN404, ARG398,
GLY403, GLU282, GLY283, and ASN279, respectively.
The interaction was done between the amino acids shown
in Fig.3, and ligand A13 was accompanied by a significant
energetic contribution to the total energy. It can be seen from
Fig.3 that there is one hydrogen bond with GLN63 at a dis-
tance of 2.64Å, three pi-pi stacked interactions with TRP
59 and TYR 62 at distances 3.77Å, 4.09Å, and 5.54Å,
respectively, and one pi-sigma interaction with TYR62 at a
distance of 4.18Å.
Table 5 Toxicity prediction for
proposed compounds Hepatotoxicity Carcinogenicity Immunotoxicity Mutagenicity Cytotoxicity LD50
A2 Inactive Inactive Inactive Active Inactive 120
A4 Active Inactive Inactive Inactive Inactive 77
A5 Active Active Active Active Inactive 96
A6 Active Active Active Active Inactive 96
A10 Inactive Inactive Inactive Active Inactive 800
A11 Active Active Active Active Inactive 675
A13 Inactive Inactive Inactive Active Inactive 800
A14 Active Active Active Active Inactive 96
Table 6 Binding affinity of
designed compounds (kcal/mol) Ligand Binding
affinity (kcal/
mol)
A2 − 7.8
A10 − 6.3
A13 − 8.1
Acarbose − 8.8
A2 A13 Acarbose
Fig. 3 The 2D and 3D interaction binding of the docking results between the protein 1HNY and inhibitors A2, A13, and acarbose
Journal of Molecular Modeling (2022) 28:106
1 3
Page 11 of 17 106
While the good binding affinity is due to the molecular
interactions implicated between A2 and the protein 1HNY.
The inhibitor A2 forms two pi-sigma bonds with TRP59 at
a distance of 3.91Å and 3.78Å, three pi-pi stacked and pi
alkyl with TYR62 at a distance of 5.96Å, 5.88Å, 4.38Å,
and 4.73Å, respectively.
The high affinity of inhibitor A13 is due to the pres-
ence of van der Waals forces (TRP58, THR163, LEU165,
ASP197, ASP300, and HIS305), which create a strong cohe-
sive environment, thereby stabilizing the complex formed.
Also, the presence of two alkyl interactions with LEU162
and ALA198 at distances of 4.18Å and 5.33Å, respectively.
Molecular dynamics simulations
The dynamic behavior of alone and its complexes in an
aqueous environment were studied using MD simulation.
The docked conformation with the lowest binding of both
ligands was used as an initial point for the MD simulation.
Analysis ofRMSD andRMSF
The preliminary assessment of MD simulation data was
done by calculating root-mean-square deviations (RMSD)
of the backbone of all systems with respect to their initial
coordinates to explore the stability of α-amylase and the
complexes. The RMSD of uncomplexed and complexed
α-amylase is shown in Fig.4A. As we can see from the data,
RMSD of (α-amylase complexed with A2 and A13) systems
were observed to be below 0.25nm during the whole simu-
lation period. Whereas, RMSD (α-amylase complexed with
acarbose) system showed a set of deviations in the Cα-atom
of protein α-amylase backbone structure along the entire
simulation timeline. The RMSD of the α-amylase-A13
complex showed some initial variations, but it became
stable after 20ns. Almost constant RMSD of the systems
(α-amylase-A2 and α-amylase-13) suggests their high sta-
bility under physiological conditions. Whereas the complex
α-amylase-acarbose did not show very convincing stability
due to the large variety of structural fluctuations known from
the α-amylase protein backbone subgroups. The fluctuation
in residues α-amylase in uncomplexed and complexed form
was determined by calculating the root-mean-square fluctua-
tion (RSF) of Cα of the protein (Fig.4B). For both systems
(α-amylase-A2 and α-amylase-13), the RMSF of most of
the amino acids of α-amylase was found to be lower than
0.15nm signifying great stability of both systems. There
were few spikes in the RMSF of both systems, which are
attributed to the random coil or loops of the protein that tend
to move in an aqueous environment. Instead, the α-amylase-
acarbose system showed high RMSF values compared to
both systems (α-amylase-A2 and α-amylase-13), which
means that the potential structural fluctuations of the Cα
atoms of the α-amylase protein are significant and there-
fore the stability of the system in the physiological environ-
ment seems to be poor. The RMSF of individual atoms of
three ligands (A2, A13, and acarbose) was also calculated
(Fig.4C). The RMSF of atoms of three ligands showed
some fluctuation, which is due to their dynamical shift at the
binding from their initial positions. The RMSD and RMSF
scores obtained indicate the greater stability of both systems
(α-amylase-A2 and α-amylase-13) compared to α-amylase-
acarbose. Thus, the molecular structures of the ligands A2
and A13 can be screened to inhibit α-amylase more stable
than acarbose. Based on these assumptions, in the follow-
ing work, we investigate the dynamics properties yielding
the potentially high stability of both systems (α-amylase-A2
and α-amylase-13).
Assessment of Rg andSASA, andenergies
The mass-weighted root-mean-square distance of a collec-
tion of atoms from their common center of mass is defined as
the radius of gyration (Rg) which is considered an important
parameter to assess the stability of proteins in an aqueous
environment [56]. In general, globular and compact proteins
Fig. 4 A Root-mean-square deviation (RMSD) of uncomplexed
α-amylase and complexed with A2, A13, and acarbose. B Root-mean-
square fluctuation (RMSF) Cα atoms of uncomplexed α-amylase and
complexed with A2, A13, and acarbose. C The average RMSF value
of each atom of A2, A13, and acarbose
Journal of Molecular Modeling (2022) 28:106
1 3
106 Page 12 of 17
show lesser variations in Rg compared to the expanded form
of the proteins [57]. The changes in Rg of α-amylase alone
and its complex with A2 and A13 are presented in Fig.5A.
The Rg of all structures were found to be roughly constant
with time showing their stability in an aqueous medium. The
average Rg of α-amylase alone, α-amylase-A2 complex, and
the α-amylase-A13 complex was found to be 2.341 ± 0.009,
2.325 ± 0.012, and 2.325 ± 0.007 nm, respectively. These
values were clearly showed that all systems were quite sta-
ble during MD simulation. Solvent accessible surface area
(SASA) of proteins is also taken into account while study-
ing the stability of proteins during MD [57, 58]. SASA of
uncomplexed α-amylase and complexed with A2 and A13
is with respect to time is presented in (Fig.5B). All sys-
tems were found to be exhibiting a constant SASA over the
entire simulation period showing their stability. The average
SASA of α-amylase alone, α-amylase-A2 complex, and the
α-amylase-A13 complex was found to be 192.848 ± 3.249,
192.476 ± 2.726, and 191.930 ± 3.701 nm2, respectively. A
negligible change in SASA of each system over simulation
time further confirms their stable nature under physiological
conditions.
Finally, the verification of stability of the systems
was also performed by calculating their physicochemical
parameters such as total and potential energies (Fig.5C).
The straight line of potential and total energy with negli-
gible fluctuations shows that all systems were well equili-
brated and remained stable during the simulation [44].
The energies of both complexes were nearly identical to
α-amylase alone.
Analysis ofthesecondary structure ofα‑amylase
andhydrogen bonds
The effect of the interaction of A2 and A13 on the second-
ary structure of α-amylase was studied by calculating the
average secondary structure from the entire trajectory of
each system (Fig.6A). The α-helix, β-sheet, and coils in
α-amylase alone were found to be 17.29, 18.41, and 23.82%,
respectively. There were negligible changes in all secondary
2
2
.
.
2
2
2
2
.
.
3
3
2
2
.
.
4
4
2
2
.
.
5
5
0
0
2
2
0
0
4
4
0
0
6
6
0
0
8
8
0
0
1
1
0
0
0
0
R
R
g
g
(
(
n
n
m
m
)
)
T
T
i
i
m
m
e
e
(
(
n
n
s
s
)
)
α
α
-
-
a
a
m
m
y
y
l
l
a
a
s
s
e
e
α
α
-
-
a
a
m
m
y
y
l
l
a
a
s
s
e
e
+
+
A
A
2
2
α
α
-
-
a
a
m
m
y
y
l
l
a
a
s
s
e
e
+
+
A
A
1
1
3
3
1
1
5
5
0
0
1
1
7
7
0
0
1
1
9
9
0
0
2
2
1
1
0
0
2
2
3
3
0
0
2
2
5
5
0
0
0
0
2
2
0
0
4
4
0
0
6
6
0
0
8
8
0
0
1
1
0
0
0
0
S
S
A
A
S
S
A
A
(
(
n
n
m
m
2
2
)
)
T
T
i
i
m
m
e
e
(
(
n
n
s
s
)
)
α
α
-
-
a
a
m
m
y
y
l
l
a
a
s
s
e
e
α
α
-
-
a
a
m
m
y
y
l
l
a
a
s
s
e
e
+
+
A
A
2
2
α
α
-
-
a
a
m
m
y
y
l
l
a
a
s
s
e
e
+
+
A
A
1
1
3
3
-1.7
-1.5
-1.3
-1.1
02040608
01
00
Energy (× 105kcal/mol)
Time (ns)
PE of α-amylase
PE of α-amylase + A2
PE of α-amylase + A13
TE of α-amylase
TE of α-amylase + A2
TE of α-amylase + A13
(A) (B) (C)
Fig. 5 A Radius of gyration (Rg) of uncomplexed α-amylase and
complexed with A2 and A13 as a function of time. B Solvent acces-
sible surface area (SASA) of uncomplexed α-amylase and complexed
with A2 and A13 as a function of time. C The potential energy and
total energy of uncomplexed α-amylase and complexed with A2 and
A13 as a function of time
0
0
5
5
1
1
0
0
1
1
5
5
2
2
0
0
2
2
5
5
3
3
0
0
P
P
e
e
r
r
c
c
e
e
n
n
t
t
a
a
g
g
e
e
S
S
e
e
c
c
o
o
n
n
d
d
a
a
r
r
y
y
s
s
t
t
r
r
u
u
c
c
t
t
u
u
r
r
e
e
o
o
f
f
p
p
r
r
o
o
t
t
e
e
i
i
n
n
α
α
-
-
a
a
m
m
y
y
l
l
a
a
s
s
e
e
α
α
-
-
a
a
m
m
y
y
l
l
a
a
s
s
e
e
+
+
A
A
2
2
α
α
-
-
a
a
m
m
y
y
l
l
a
a
s
s
e
e
+
+
A
A
1
1
3
3
0
0
1
1
2
2
3
3
4
4
5
5
0
0
2
2
0
0
4
4
0
0
6
6
0
0
8
8
0
0
1
1
0
0
0
0
H
H
y
y
d
d
r
r
o
o
g
g
e
e
n
n
B
B
o
o
n
n
d
d
s
s
(
(
N
N
u
u
m
m
b
b
e
e
r
r
)
)
T
T
i
i
m
m
e
e
(
(
n
n
s
s
)
)
α
α
-
-
a
a
m
m
y
y
l
l
a
a
s
s
e
e
+
+
A
A
2
2
α
α
-
-
a
a
m
m
y
y
l
l
a
a
s
s
e
e
+
+
A
A
1
1
3
3
(A) (B)
Fig. 6 A Percentage of secondary structure in uncomplexed α-amylase and complexed with A2 and A13. B Number of hydrogen bonds of A2
and A13 with α-amylase
Journal of Molecular Modeling (2022) 28:106
1 3
Page 13 of 17 106
structural components of α-amylase by the interaction of
both ligands (A2 and A13). The results clearly show that
the interaction of both ligands did not alter the secondary
structure of the protein.
The binding of A2 and A13 with α-amylase was further
explored by calculating the hydrogens bond profiles and the
number of hydrogen bonds during MD simulation (Fig.6B).
The average number of hydrogen bonds in 10,000 frames
of the α-amylase-A2 complex was slightly higher than the
α-amylase-A13 complex. The existence of hydrogen bonds
in the entire trajectory of both complexes was also deter-
mined. The existence of hydrogen bonds was found in the
hydrogen bond profile of the complexes. He changes in
hydrogen bond profiles show the dynamics shift of ligand
in the binding pocket and changes in nature of interactions
occurring.
Calculation ofbinding energies andidentification ofkey
residues involved inbinding
The detailed insight regarding the binding ener-
gies involved in the interaction of A2 and A13 with
α-amylase was performed using MM-PBSA calcula-
tions. Typically, in ligand–protein interactions, the
predominant forces involved are non-covalent interac-
tions. The major forces are electrostatic, hydrophobic
interactions, hydrogen bonds, van der Waals force, etc.
These binding forces may either contribute positively
or negatively to the overall binding of the ligand with
protein [59]. The MM-PBSA binding energies were cal-
culated by extracting 50 frames from 50 to 100ns of the
trajectories at 1ns interval (Table7). The major con-
tribution in the total binding of both ligands was elec-
trostatic energy and van der Waals energy. There was
also a small contribution of SASA energy. However,
polar solvation energy impaired the interaction of both
ligands with α-amylase. The overall binding energy for
the interaction of A2 and A13 with α-amylase was found
to be − 50.36 ± 0.97 and − 60.52 ± 0.54 kcal mol−1,
respectively.
Using MM-PBSA calculations, the binding energies
of individual residues were also calculated (Table8).
Glu18, Glu60, Asp96, Asp147, Glu149, Asp153, Asp167,
Glu233, Asp236, Glu240, Glu255, Asp300, Asp353, and
Asp356 are major contributors for the interaction of
A2 with α-amylase. Similarly, Glu18, Glu60, Asp147,
Glu149, Asp167, Asp197, Glu233, Asp300, Asp353, and
Asp356 were found to be the largest energy contributors
for A13-α-amylase interaction.
Table 7 Binding free energy (kcal mol-1) for the interaction of A2
and A13 with α-amylase using MMBSA analysis
ΔEvdW: van der Waal energy, ΔEele: electrostatic energy, ΔEPSE: polar
solvation energy, ΔESASA: solvent accessible surface area energy,
ΔEBE: binding energy
Type of energy Ligands
A2 A13
ΔEvdW − 35.31 ± 0.68 − 33.18 ± 0.33
ΔEele − 58.45 ± 0.70 − 50.16 ± 0.53
ΔEPSE 48.01 ± 1.75 26.85 ± 0.62
ΔESSASA − 4.62 ± 0.05 − 4.02 ± 0.03
ΔEBE − 50.36 ± 0.97 − 60.52 ± 0.54
Table 8 The average polar,
apolar, and total binding
energies (kcal mol−1) of the key
residues
Epolar: polar energy; EApolar: apolar energy; Etotal: total energy
A2 A13
Residues Epolar EApolar Etotal Residues Epolar EApolar Etotal
Glu18 0.729 0.000 − 4.001 GLU-18 1.651 0.000 − 4.176
Glu60 0.609 0.000 − 4.052 GLU-60 2.155 0.000 − 4.798
Asp96 1.909 0.000 − 4.382 ASP-147 1.227 0.000 − 5.622
Asp147 1.069 0.000 − 5.127 GLU-149 0.237 0.000 − 4.550
Glu149 0.674 -0.001 − 5.125 ASP-167 1.261 0.000 − 4.089
Asp153 1.152 0.000 − 4.532 ASP-197 2.543 − 0.003 − 5.515
Asp167 1.130 0.000 − 4.149 GLU-233 1.541 0.000 − 5.043
Glu233 6.288 -0.037 − 5.485 ASP-300 3.024 − 0.052 − 5.686
Asp236 0.872 0.000 − 4.677 ASP-353 0.611 0.000 − 4.528
Glu240 1.307 0.000 − 4.680 ASP-356 1.801 − 0.002 − 5.769
Glu255 0.679 0.000 − 4.141
Asp300 4.926 -0.043 − 5.457
Asp353 0.176 0.000 − 4.037
Asp356 0.405 -0.000 − 5.182
Journal of Molecular Modeling (2022) 28:106
1 3
106 Page 14 of 17
Principal component analysis
Principal component analysis (PCA) is a standard statistical
method to study the large-scale motion in proteins which
is performed by reducing the dimensionality of the data
set without losing important information, which is charac-
terized by eigenvectors [60]. PCA was perfumed to deci-
pher the differences in flexibility parameters between the
α-amylase and both complexes (Fig.7A). Both the com-
plexes occupied a slightly larger conformational space com-
pared to α-amylase alone indicating slightly more structural
stability of the complexes than the protein alone. The free
energy landscapes of all systems were plotted to decipher
the variations in the respective protein folding patterns
(Fig.8). It is evident from the landscape that all systems
reached energy minima however, the changes in the position
of energy minima denotes a slight alteration in the confor-
mation of α-amylase caused by the interaction of ligands.
The lowest energy minima structures were extracted for the
respective trajectories and then Ramachandran plots were
made for the energy minima structures (Fig.7B). The aver-
age phi (φ) and psi (ψ) angles for α-amylase alone were
59.54 and 89.99, respectively. Similarly, φ and ψ angles for
α-amylase-A2 complex and α-amylase-A13 complex were
found to be 60.82 and 91.37 and 60.22 and 89.78, respec-
tively. A slight variation in the dihedral angles with respect
-6
-4
-2
0
2
4
6
-8 -6 -4 -2 0246810
projection on eigenvector 2 (nm)
projection on eigenvector 1
α-amylase
α-amylase + A2
α-amylase + A13
-180
0
180
-180 0 180
Psi
Phi
α-amylase
α-amylase + A2
α-amylase + A13
(A) (B)
PCA 2D proj
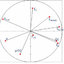
Fig. 7 PCA of uncomplexed α-amylase and complexed with A2 and A13. B Ramachandran plot of the energy minima of uncomplexed
α-amylase and complexed with A2 and A13
A13protein
A2
(C)(A) (B)
Fig. 8 Free energy landscape plot of A α-amylase alone, B α-amylase-A2 complex, and C α-amylase-A13 complex
Journal of Molecular Modeling (2022) 28:106
1 3
Page 15 of 17 106
to the protein alone shows the dynamical shift in the struc-
tural conformation of α-amylase ligands.
Conclusion
In this study, QSAR was used to develop MLR models
capable of discovering new molecules that may be used
as alpha-amylase inhibitors. The obtained models were
examined both internally and externally to determine their
statistical quality.
We chose the best model and used it to design new
molecules with good activity against alpha-amylase.
The pharmacokinetic properties of the designed mol-
ecules were analyzed using ADMET, and only three
molecules with a low toxic effect were kept.
We used molecular docking to determine the best poses
for these molecules. The dynamic molecular test is the final
step that it used to determine the stability of these molecules.
As a result, these findings can be used in the development of
new compounds with the desired activity.
Supplementary Information The online version contains supplemen-
tary material available at https:// doi. org/ 10. 1007/ s00894- 022- 05097-9.
Author contribution Oussama Abchir: conceived and designed the
experiments; performed the experiments; contributed reagents, mate-
rials, analysis tools, or data; and wrote the paper. Mebarka Ouassaf, and
Faizan Abul Qais: conceived and designed the experiments; analyzed
and interpreted the data; contributed reagents, materials, analysis tools,
or data; and wrote the paper. Salah Belaidi and Said Belaaouad: con-
ceived and designed the experiments. Ossama Daoui and Souad Elkhat-
tabi: analyzed and interpreted the data and revised the paper. Samir
Chtita:conceived and designed the experiments; analyzed and inter-
effects of diabetes on your bodpreted the data; contributed reagents,
materials, analysis tools, or data; wrote the paper andsupervision.
Data Availability Not applicable.
Code availability Not applicable.
Declarations
Competing interests The authors declare no competing interests.
References
1. Danaei G, Lawes CM, Vander Hoorn S, Murray CJ, Ezzati M
(2006) Global and regional mortality from ischaemic heart disease
and stroke attributable to higher-than-optimum blood glucose con-
centration: comparative risk assessment. Lancet 368:1651–1659
2. The effects of diabetes on your body, Healthline (2014)https://
www. healt hline. com/ health/ diabe tes/ effec ts- on- body. Accessed
26 Mar2022
3. Sarwar N, Gao P, Seshasai SR, Gobin R, Kaptoge S, Di Angelan-
tonio E, Ingelsson E, Lawlor DA, Selvin E, Stampfer M (2010)
Emerging risk factors collaboration diabetes mellitus, fasting
blood glucose concentration, and risk of vascular disease: a
collaborative meta-analysis of 102 prospective studies. Lancet
375:2215–2222
4. Chun BY, Rizzo JF III (2016) Dominant optic atrophy: updates
on the pathophysiology and clinical manifestations of the optic
atrophy 1 mutation. Curr Opin Ophthalmol 27:475–480
5. Saran R, Robinson B, Abbott KC, Agodoa LY, Albertus P, Aya-
nian J, Balkrishnan R, Bragg-Gresham J, Cao J, Chen JL (2017)
US renal data system 2016 annual data report: epidemiology of
kidney disease in the United States. Am J Kidney Dis 69:A7–A8
6. Diabetes mellitus guide: causes, symptoms and treatment options
(n.d.)https:// www . drugs. com/ health- guide/ diabe tes- melli tus. html.
Accessed 26 Mar2022
7. Funke I, Melzig MF (2006) Plantas tradicionalmente utilizadas na
terapia da diabetes: fitomedicamentos como inibidores da ativi-
dade alfa-amilase, Revista Brasileira de. Farmacognosia 16:1–5
8. Sales PM, Souza PM, Simeoni LA, Magalhães PO, Silveira D
(2012) α-Amylase inhibitors: a review of raw material and isolated
compounds from plant source. J Pharm Pharm Sci 15:141–183
9. Van De Laar FA, Lucassen PL, Akkermans RP, Van De Lisdonk
EH, Rutten GE, Van Weel C (2005) α-Glucosidase inhibitors for
patients with type 2 diabetes: results from a Cochrane systematic
review and meta-analysis. Diabetes Care 28:154–163
10. Arshad T, Khan KM, Rasool N, Salar U, Hussain S, Tahir T,
Ashraf M, Waddod A, Riaz M, Perveen S, Taha M, Ismail NH
(2016) Syntheses, invitro evaluation and molecular docking
studies of 5-bromo-2-aryl benzimidazoles as α-glucosidase
inhibitors. Med Chem Res 25:2058–2069. https:// doi. org/ 10. 1007/
s00044- 016- 1614-y
11. Aminpour M, Montemagno C, Tuszynski JA (2019) An overview
of molecular modeling for drug discovery with specific illustrative
examples of applications. Molecules 24:1693
12. Chalkha M, Akhazzane M, Moussaid FZ, Daoui O, Nakkabi A,
Bakhouch M, Chtita S, Elkhattabi S, Housseini AI, Yazidi ME
(2022) Design, synthesis, characterization, invitro screening,
molecular docking, 3D-QSAR, and ADME-Tox investigations
of novel pyrazole derivatives as antimicrobial agents. New J
Chem.46:2747–2760https:// doi. org/ 10. 1039/ D1NJ0 5621B
13. Adegboye AA, Khan KM, Salar U, Aboaba SA, Chigurupati S,
Fatima I, Taha M, Wadood A, Mohammad JI, Khan H (2018)
2-Aryl benzimidazoles: synthesis, invitro α-amylase inhibi-
tory activity, and molecular docking study. Eur J Med Chem
150:248–260
14. ChemOffice, PerkinElmer Informatics.http:// www. cambr idges oft.
com, 2016. Accessed 26 Mar2022
15. MarvinSketch 5.11.4., Chem Axon 2012. https:// chema xon. com/
produ cts/ marvin. Accessed 26 Mar2022
16. ACDLABS 10. Advanced Chemistry Development. Inc. Toronto.
ON. Canada 2015. Available:(http:// www. acdla bs. com/ resso urces/
freew are/ chems ketch/). Accessed 26 Mar2022
17. XLSTAT, Software, XLSTAT company.www. xlstat. com, 2013.
Accessed 26 Mar2022
18. Chtita S, Ghamali M, Ousaa A, Aouidate A, Belhassan A, Taourati
AI, Masand VH, Bouachrine M, Lakhlifi T (2019) QSAR study
of anti-Human African trypanosomiasis activity for 2-phenylimi-
dazopyridines derivatives using DFT and Lipinski’s descriptors.
Heliyon 5(3):e01304
19. Chtita S, Aouidate A, Belhassan A, Ousaa A, Taourati AI, Elid-
rissi B, Ghamali M, Bouachrine M, Lakhlifi T (2020) QSAR study
of N-substituted oseltamivir derivatives as potent avian influenza
virus H5N1 inhibitors using quantum chemical descriptors and
statistical methods. New J Chem 44:1747–1760
20. Elidrissi B, Ousaa A, Ghamali M, Chtita S, Ajana MA, Bouachrine
M, Lakhlifi T (2014) Toxicity in vivo of nitro-aromatic
Journal of Molecular Modeling (2022) 28:106
1 3
106 Page 16 of 17
compounds: DFT and QSAR results. J Comput Aided Mol Des
4:28–37
21. Daoui O, Elkhattabi S, Chtita S, Elkhalabi R, Zgou H, Benjelloun
AT (2021) QSAR, Molecular Docking and ADMET properties
in silico studies of Novel 4, 5, 6, 7-Tetrahydrobenzo [D]-Thiazol-
2-Yl Derivatives Derived from Dimedone as potent Anti-Tumor
agents through inhibition of C-Met receptor Tyrosine Kinase.
Heliyon 7(7):E07463. https:// doi. org/ 10. 1016/j. heliy on. 2021.
e07463
22. Ouassaf M, Belaidi S, Benbrahim I, Belaidi H, Chtita S (2020)
Quantitative structure-activity relationships of 1.2. 3 triazole
derivatives as aromatase inhibition activity, Turkish Computa-
tional and Theoretical Chemistry 4(1):1–11. https:// doi. org/ 10.
33435/ tcand tc. 545369
23. Chtita S, Ghamali M, Hmamouchi R, Elidrissi B, Bourass M, Larif
M, Bouachrine M, Lakhlifi T (2016) Investigation of antileish-
manial activities of acridines derivatives against promastigotes
and amastigotes form of parasites using quantitative structure
activity relationship analysis, Advances in Physical Chemistry
e5137289.https:// doi. org/ 10. 1155/ 2016/ 51372 89
24. Diana G, Tommasi C (2002) Cross-validation methods in prin-
cipal component analysis: a comparison. Stat Methods Appl
11:71–82
25. Chatterjee S, Hadi AS, Regression analysis by example, John
Wiley & Sons, 2013
26. Rücker C, Rücker G, Meringer M (2007) Y-randomization–a
useful tool in QSAR validation, or folklore. J Chem Inf Model
47:2345–2357
27. Eriksson L, Jaworska J, Worth AP, Cronin MT, McDowell RM,
Gramatica P (2003) Methods for reliability and uncertainty assess-
ment and for applicability evaluations of classification-and regres-
sion-based QSARs. Environ Health Perspect 111:1361–1375
28. Netzeva TI, Worth AP, Aldenberg T, Benigni R, Cronin MT, Gra-
matica P, Jaworska JS, Kahn S, Klopman G, Marchant CA (2005)
Current status of methods for defining the applicability domain
of (quantitative) structure-activity relationships: the report and
recommendations of ECVAM workshop 52. Altern Lab Anim
33:155–173
29. Sabando MV, Ponzoni I, Soto AJ (2019) Neural-based approaches
to overcome feature selection and applicability domain in drug-
related property prediction. Appl Soft Comput 85:105777
30. Jaworska J, Nikolova-Jeliazkova N, Aldenberg T (2005) QSAR
applicability domain estimation by projection of the training set
in descriptor space: a review. Altern Lab Anim 33:445–459
31. Zoete V, Daina A, Bovigny C, Michielin O (2016) SwissSimi-
larity: a web tool for low to ultra high throughput ligand-based
virtual screening
32. Daina A, Michielin O, Zoete V (2017) SwissADME: a free web
tool to evaluate pharmacokinetics, drug-likeness and medicinal
chemistry friendliness of small molecules. Sci Rep 7:1–13
33. U.N.E.C. for E. Secretariat, United Nations. Economic Commis-
sion for Europe. Secretariat. Globally Harmonized System of
Classification and Labelling of Chemicals (GHS)., United Nations
Publications, 2013.
34. Daoui O, Mazoir N, Bakhouch M, etal (2022) 3D-QSAR, ADME-
Tox, and molecular docking of semisynthetic triterpene deriva-
tives as antibacterial and insecticide agents. Struct Chem. https://
doi. org/ 10. 1007/ s11224- 022- 01912-4
35. Shou WZ (2020) Current status and future directions of high-
throughput ADME screening in drug discovery. Journal of Phar-
maceutical Analysis 10:201–208
36. Chtita S, Belhassan A, Aouidate A, Belaidi S, Bouachrine M,
Lakhlifi T (2021) Discovery of potent SARS-CoV-2 inhibitors
from approved antiviral drugs via docking and virtual screening.
Comb Chem High Throughput Screening 24:441–454
37. Ouassaf M, Belaidi S, Mogren Al Morgen M, Chtita S, Ullah
Khan S, ThetHtar T (2021) Combined docking methods and
molecular dynamics to identify effective antiviral 2, 5-diamin-
obenzophenonederivatives against SARS-CoV-2. J King Saud
Univ Sci 33:101352. https:// doi. org/ 10. 1016/j. jksus. 2021. 101352
38. Brayer GD, Luo Y, Withers SG (1995) The structure of human
pancreatic α-amylase at 1.8 Å resolution and comparisons with
related enzymes. Protein Sci 4:1730–1742
39. Sybyl-X 2.0, Tripos International, St. Louis, Missouri, 63144,
USA., Software Informer. (n.d.).https:// sybyl-x. softw are. infor mer.
com/2. 0/. Accessed 26 Mar2022
40. Morris GM, Huey R, Lindstrom W, Sanner MF, Belew RK, Good-
sell DS, Olson AJ (2009) AutoDock4 and AutoDockTools4: auto-
mated docking with selective receptor flexibility. J Comput Chem
30:2785–2791
41. Trott O, Olson AJ (2010) AutoDock Vina: improving the speed
and accuracy of docking with a new scoring function, efficient
optimization, and multithreading. J Comput Chem 31:455–461
42. Dassault Systèmes BIOVIA Discovery Studio Modeling Environ-
ment. Release 2017 Dassault Systèmes 2016., (n.d.). https:// disco
ver. 3ds. com/ disco very- studio- visua lizer- downl oad. Accessed 26
Mar2022
43. H.J.C. Berendsen, B. Hess, E. Lindahl, D. Van Der Spoel, A.E.
Mark, G. Groenhof, van der Spoel D, Lindahl E, Hess B, Groen-
hof G, Mark AE, Berendsen HJC. (2005) GROMACS: fast, flex-
ible, and free. J of Comput Chem 26(16):1701–1718. https:// doi.
org/ 10. 1002/ jcc. 20291
44. Liao S-Y, Mo G-Q, Chen J-C, Zheng K-C (2014) Exploration
of the binding mode between (-)-zampanolide and tubulin using
docking and molecular dynamics simulation. J Mol Model 20:1–9
45. Sousa da Silva AW, Vranken WF (2012) ACPYPE-Antechamber
python parser interface. BMC Res Notes 5:1–8
46. Brand W, Oosterhuis B, Krajcsi P, Barron D, Dionisi F, Van
Bladeren PJ, Rietjens IM, Williamson G (2011) Interaction of
hesperetin glucuronide conjugates with human BCRP, MRP2 and
MRP3 as detected in membrane vesicles of overexpressing bacu-
lovirus-infected Sf9 cells. Biopharm Drug Dispos 32:530–535
47. Parrinello M, Rahman A, (2002) Polymorphic transítions in single
crystals: a new molecular dynamics method polymorphic transí-
tions in single crystals: a new molecular dynamics method
48. Kumari R., Kumar C, (2014) Open source drug discovery and A
Lynn J Chem Inf Model 54:10–1021
49. Dahmani R, Manachou M, Belaidi S, Chtita S, Boughdiri S (2021)
Structural characterization and QSAR modeling of 1,2,4-triazole
derivatives as α-glucosidase inhibitors. New J Chem 45:1253–
1261. https:// doi. org/ 10. 1039/ D0NJ0 5298A
50. Chtita S, Belhassan A, Bakhouch M, Taourati AI, Aouidate A,
Belaidi S, Moutaabbid M, Belaaouad S, Bouachrine M, Lakhlifi
T (2021) QSAR study of unsymmetrical aromatic disulfides as
potent avian SARS-CoV main protease inhibitors using quantum
chemical descriptors and statistical methods. Chemom Intell Lab
Syst 210:104266
51. Lipinski CA, Lombardo F, Dominy BW, Feeney PJ (1997) Experi-
mental and computational approaches to estimate solubility and
permeability in drug discovery and development settings. Adv
Drug Deliv Rev 23:3–25
52. Veber DF, Johnson SR, Cheng H-Y, Smith BR, Ward KW, Kopple
KD (2002) Molecular properties that influence the oral bioavail-
ability of drug candidates. J Med Chem 45:2615–2623
53. Ghose AK, Viswanadhan VN, Wendoloski JJ (1999) A knowl-
edge-based approach in designing combinatorial or medicinal
chemistry libraries for drug discovery. 1. A qualitative and quan-
titative characterization of known drug databases. J Comb Chem
1:55–68
54. Szakács G, Váradi A, Özvegy-Laczka C, Sarkadi B (2008)
The role of ABC transporters in drug absorption, distribution,
Journal of Molecular Modeling (2022) 28:106
1 3
Page 17 of 17 106
metabolism, excretion and toxicity (ADME–Tox). Drug Discovery
Today 13:379–393
55. Hollenberg PF (2002) Characteristics and common properties of
inhibitors, inducers, and activators of CYP enzymes. Drug Metab
Rev 34:17–35
56. Qais FA, Sarwar T, Ahmad I, Khan RA, Shahzad SA, Husain FM
(2021) Glyburide inhibits non-enzymatic glycation of HSA: an
approach for the management of AGEs associated diabetic com-
plications. Int J Biol Macromol 169:143–152
57. Fouedjou RT, Chtita S, Bakhouch M, Belaidi S, Ouassaf M,
Djoumbissie LA, Tapondjou LA, Qais FA (2021) Cameroonian
medicinal plants as potential candidates of SARS-CoV-2 inhibi-
tors. J Biomol Struct Dyn 0:1–15. https:// doi. org/ 10. 1080/ 07391
102. 2021. 19141 70
58. Rath B, Qais FA, Patro R, Mohapatra S, Sharma T (2021) Design,
synthesis and molecular modeling studies of novel mesalamine
linked coumarin for treatment of inflammatory bowel disease.
Bioorganic Med Chem Lett 41:128029
59. Siddiqui S, Ameen F, Jahan I, Nayeem SM, Tabish M (2019) A
comprehensive spectroscopic and computational investigation on
the binding of the anti-asthmatic drug triamcinolone with serum
albumin. New J Chem 43:4137–4151
60. Siddiqui S, Ameen F, Kausar T, Nayeem SM, Rehman SU,
Tabish M (2021) Biophysical insight into the binding mecha-
nism of doxofylline to bovine serum albumin: an invitro and in
silico approach. Spectrochim Acta Part A: Mol Biomol Spectrosc
249:119296
Publisher's Note Springer Nature remains neutral with regard to
jurisdictional claims in published maps and institutional affiliations.