Figure - available from: Molecular Psychiatry
This content is subject to copyright. Terms and conditions apply.
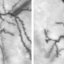
Taok2 deficiency affects cortex morphology a Taok2 downregulation changes the position of upper-layer neurons at P7. Both brain hemispheres were electroporated in utero at E15 with control shRNA (mCherry) and Taok2 shRNA (Venus) on the contralateral sides. b Quantification of cell distribution at P7 (****p < 0.0001 by unpaired t-test; n = 6–7 brains per condition; median is represented by black line; see Supplementary Fig. 10a for individual brains). c Taok2 KO neurons, labeled in utero at E15, are localized more superficially in the upper cortex at postnatal day 21 than WT or Het neurons. d Quantification of cell distribution at P21 (p < 0.0001 by one-way ANOVA, post hoc Dunnett’s multiple test ****p < 0.0001; n = 3 brains per condition; median is represented by black line; see Supplementary Fig. 10b for individual brains). e Longitudinal MRI imaging of brains from WT and Taok2 KO mice shows morphological changes in the cortex in Taok2 KO brains compared with WT brains. f Quantification of cortex volume (upper panel) and cortex curvature (lower panel) at different time points (p < 0.0001 by one-way ANOVA, post hoc Dunnett’s test **p < 0.01, ***p < 0.001 and ****p < 0.0001, WT = 6–8 mice, Het = 9–15 mice, KO = 7–12 mice; values are the mean ± s.e.m.).
Source publication
The precise development of the neocortex is a prerequisite for higher cognitive and associative functions. Despite numerous advances that have been made in understanding neuronal differentiation and cortex development, our knowledge regarding the impact of specific genes associated with neurodevelopmental disorders on these processes is still limit...
Similar publications
Disordered cellular development, abnormal neuroanatomical formations, and dysfunction of neuronal circuitry are among the pathological manifestations of cortical regions in the brain that are often implicated in complex neurodevelopmental disorders. With the advancement of stem cell methodologies such as cerebral organoid generation, it is possible...
The mosaic variegated aneuploidy (MVA)-associated gene Budding Uninhibited by Benzimidazole 1B (BUB1B) encodes BUBR1, a core member of the spindle assembly checkpoint complex that ensures kinetochore-spindle attachment for faithful chromosome segregation. BUB1B mutation in humans and its deletion in mice cause microcephaly. In the absence of BubR1...
Idiopathic autism spectrum disorder (ASD) is highly heterogeneous, and it remains unclear how convergent biological processes in affected individuals may give rise to symptoms. Here, using cortical organoids and single-cell transcriptomics, we modeled alterations in the forebrain development between boys with idiopathic ASD and their unaffected fat...
Development of the mammalian neocortex requires proper inside‐out migration of developing cortical neurons from the germinal ventricular zone toward the cortical plate. The mechanics of this migration requires precise coordination of different cellular phenomena including cytoskeleton dynamics, membrane trafficking, and cell adhesion. The small GTP...
We analyze more than 700,000 single-nucleus RNA-seq profiles from 106 donors during prenatal and postnatal developmental stages and identify lineage-specific programs that underlie the development of specific subtypes of excitatory cortical neurons, interneurons, glial cell types and brain vasculature. By leveraging single-nucleus chromatin accessi...
Citations
... Previously, we and others reported that, while TAOK2α is associated with microtubules (23,27), TAOK2β has a functional association with the actin cytoskeleton through binding to and affecting RhoA activity (25). However, the role of TAOK2 in neuronal differentiation is still not well understood. ...
Genes implicated in translation control have been associated with autism spectrum disorders (ASDs). However, some important genetic causes of autism, including the 16p11.2 microdeletion, bear no obvious connection to translation. Here, we use proteomics, genetics, and translation assays in cultured cells and mouse brain to reveal altered translation mediated by loss of the kinase TAOK2 in 16p11.2 deletion models. We show that TAOK2 associates with the translational machinery and functions as a translational brake by phosphorylating eukaryotic elongation factor 2 (eEF2). Previously, all signal-mediated regulation of translation elongation via eEF2 phosphorylation was believed to be mediated by a single kinase, eEF2K. However, we show that TAOK2 can directly phosphorylate eEF2 on the same regulatory site, but functions independently of eEF2K signaling. Collectively, our results reveal an eEF2K-independent signaling pathway for control of translation elongation and suggest altered translation as a molecular component in the etiology of some forms of ASD.
... Our laboratory is now embracing these advancements by processing brains in 3D using block face serial imaging. While the voxel resolution in this method is ten times higher than what current MRI technologies offer for rodent brains [23], it is clear that much of the groundwork in automated segmentation will likely stem from these studies since they have long established that manually segmenting 3D volumes poses even greater challenges. ...
Using a high-throughput neuroanatomical screen of histological brain sections developed in collaboration with the International Mouse Phenotyping Consortium, we previously reported a list of 198 genes whose inactivation leads to neuroanatomical phenotypes. To achieve this milestone, tens of thousands of hours of manual image segmentation were necessary. The present work involved developing a full pipeline to automate the application of deep learning methods for the automated segmentation of 24 anatomical regions used in the aforementioned screen. The dataset includes 2000 annotated parasagittal slides (24,000 × 14,000 pixels). Our approach consists of three main parts: the conversion of images (.ROI to .PNG), the training of the deep learning approach on the compressed images (512 × 256 and 2048 × 1024 pixels of the deep learning approach) to extract the regions of interest using either the U-Net or Attention U-Net architectures, and finally the transformation of the identified regions (.PNG to .ROI), enabling visualization and editing within the Fiji/ImageJ 1.54 software environment. With an image resolution of 2048 × 1024, the Attention U-Net provided the best results with an overall Dice Similarity Coefficient (DSC) of 0.90 ± 0.01 for all 24 regions. Using one command line, the end-user is now able to pre-analyze images automatically, then runs the existing analytical pipeline made of ImageJ macros to validate the automatically generated regions of interest resulting. Even for regions with low DSC, expert neuroanatomists rarely correct the results. We estimate a time savings of 6 to 10 times.
... More recently, cohort-based sequencing studies have provided a genetic framework to studying ASD and have identified several hundred implicated genes. Genetic disruptions include inherited rare variants and less common de novo mutations that exist as single nucleotide polymorphisms (SNPs), copy number variants (CNVs), and chromosomal abnormalities [12][13][14][15][16][17][18][19] Despite the immense progress in identifying ASD-risk genes, the encoded proteins and resulting pathobiology remains elusive. Scientists have turned to genetic modeling to better understand the molecular, cellular, and functional (circuit-based) consequences to disruption in these ASD-risk genes [20]. ...
Autism spectrum disorder (ASD) is a complex neurodevelopmental disorder caused by genetic or environmental perturbations during early development. Diagnoses are dependent on the identification of behavioral abnormalities that likely emerge well after the disorder is established, leaving critical developmental windows uncharacterized. This is further complicated by the incredible clinical and genetic heterogeneity of the disorder that is not captured in most mammalian models. In recent years, advancements in stem cell technology have created the opportunity to model ASD in a human context through the use of pluripotent stem cells (hPSCs), which can be used to generate 2D cellular models as well as 3D unguided- and region-specific neural organoids. These models produce profoundly intricate systems, capable of modeling the developing brain spatiotemporally to reproduce key developmental milestones throughout early development. When complemented with multi-omics, genome editing, and electrophysiology analysis, they can be used as a powerful tool to profile the neurobiological mechanisms underlying this complex disorder. In this review, we will explore the recent advancements in hPSC-based modeling, discuss present and future applications of the model to ASD research, and finally consider the limitations and future directions within the field to make this system more robust and broadly applicable.
... This effect could be replicated in TAOK2 KO brains but also in the heterozygous 16p11.2 DEL animals, that displayed reduced levels of phosphorylated JNK1 and neuronal migration deficits, that could be partially rescued by ectopic expression of TAOK2α in in the developing cortex (Scharrenberg et al., 2022). ...
Neurodevelopmental disorders (NDDs) include a broad spectrum of pathological conditions that affect >4% of children worldwide, share common features and present a variegated genetic origin. They include clinically defined diseases, such as autism spectrum disorders (ASD), attention-deficit/hyperactivity disorder (ADHD), motor disorders such as Tics and Tourette’s syndromes, but also much more heterogeneous conditions like intellectual disability (ID) and epilepsy. Schizophrenia (SCZ) has also recently been proposed to belong to NDDs. Relatively common causes of NDDs are copy number variations (CNVs), characterised by the gain or the loss of a portion of a chromosome. In this review, we focus on deletions and duplications at the 16p11.2 chromosomal region, associated with NDDs, ID, ASD but also epilepsy and SCZ. Some of the core phenotypes presented by human carriers could be recapitulated in animal and cellular models, which also highlighted prominent neurophysiological and signalling alterations underpinning 16p11.2 CNVs-associated phenotypes. In this review, we also provide an overview of the genes within the 16p11.2 locus, including those with partially known or unknown function as well as non-coding RNAs. A particularly interesting interplay was observed between MVP and MAPK3 in modulating some of the pathological phenotypes associated with the 16p11.2 deletion. Elucidating their role in intracellular signalling and their functional links will be a key step to devise novel therapeutic strategies for 16p11.2 CNVs-related syndromes.
Neurodevelopmental disorders (NDDs) are polygenic in nature and copy number variants (CNVs) are ideal candidates to study the nature of this polygenic risk. The disruption of striatal circuits is considered a central mechanism in NDDs. The 16p11.2 hemi-deletion (16p11.2 del/+) is one of the most common CNVs associated with NDD, and 16p11.2 del/+ mice show sex-specific striatum-related behavioral phenotypes. However, the critical genes among the 27 genes in the 16p11.2 region that underlie these phenotypes remain unknown. Previously, we applied a novel strategy to identify candidate genes associated with the sex-specific phenotypes of 16p11.2 del/+ mice and highlighted three genes within the deleted region: thousand and one amino acid protein kinase 2 (Taok2), seizure-related 6 homolog-like 2 (Sez6l2), and major vault protein (Mvp). Using CRISPR/Cas9, we generated mice carrying null mutations in Taok2, Sez6l2, and Mvp (3 gene hemi-deletion (3g del/+)). Hemi-deletion of these 3 genes recapitulates sex-specific behavioral alterations in striatum-dependent behavioral tasks observed in 16p11.2 del/+ mice, specifically male-specific hyperactivity and impaired motivation for reward seeking. Moreover, RNAseq analysis revealed that 3g del/+ mice exhibit gene expression changes in the striatum similar to 16p11.2 del/+ mice exclusively in males. Subsequent analysis identified translation dysregulation and/or extracellular signal-regulated kinase signaling as plausible molecular mechanisms underlying male-specific, striatum-dependent behavioral alterations. Interestingly, ribosomal profiling supported the notion of translation dysregulation in both 3g del/+ and 16p11.2 del/+ male mice. However, mice carrying a 4-gene deletion (with an additional deletion of Mapk3) exhibited fewer phenotypic similarities with 16p11.2 del/+ mice. Together, the mutation of 3 genes within the 16p11.2 region phenocopies striatal sex-specific phenotypes of 16p11.2 del/+ mice. These results support the importance of a polygenic approach to study NDDs and underscore that the effects of the large genetic deletions result from complex interactions between multiple candidate genes.
Thousand and one amino acid kinases (TAOKs) are relatively understudied and functionally pleiotropic protein kinases that have emerged as important regulators of neurodevelopment. Through their conserved amino-terminal catalytic domain, TAOKs mediate phosphorylation at serine/threonine residues in their substrates, but it is their divergent regulatory carboxyl-terminal domains that confer both exquisite functional specification and cellular localization. In this Review, we discuss the physiological roles of TAOKs and the intricate signaling pathways, molecular interactions, and cellular behaviors they modulate—from cell stress responses, division, and motility to tissue homeostasis, immunity, and neurodevelopment. These insights are then integrated into an analysis of the known and potential impacts of disease-associated variants of TAOKs, with a focus on neurodevelopmental disorders, pain and addiction, and neurodegenerative diseases. Translating this foundation into clinical benefits for patients will require greater structural and functional differentiation of the TAOKs afforded by their individually specialized domains.
Copy number variants (CNVs) are genomic imbalances strongly linked to the aetiology of neuropsychiatric disorders such as schizophrenia and autism. By virtue of their large size, CNVs often contain many genes, providing a multi-genic view of disease processes that can be dissected in model systems. Thus, CNV research provides an important stepping stone towards understanding polygenic disease mechanisms, positioned between monogenic and polygenic risk models. In this review, we will outline hypothetical models for gene interactions occurring within CNVs and discuss different approaches used to study rodent and stem cell disease models. We highlight recent work showing that genetic and pharmacological strategies can be used to rescue important aspects of CNV-mediated pathophysiology, which often converges onto synaptic pathways. We propose that using a rescue approach in complete CNV models provides a new path forward for precise mechanistic understanding of complex disorders and a tangible route towards therapeutic development.